| Signature Domain? help Back to Top |
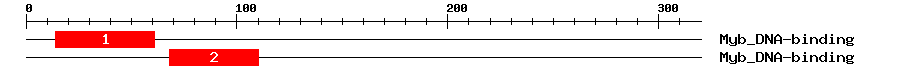 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | Myb_DNA-binding | 61.2 | 2.1e-19 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg WT+eEd +l+ +++q+G+++W++I++ g+ R++k+c++rw +yl
AT5G56110.1 14 RGQWTPEEDNKLASYIAQHGTRNWRLIPKNAGLQRCGKSCRLRWTNYL 61
89********************************************97 PP
|
| 2 | Myb_DNA-binding | 45.7 | 1.6e-14 | 68 | 110 | 2 | 46 |
SSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 2 grWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
g +++ E+ ++v+ + lG++ W++Ia+ ++ gRt++++k++w++
AT5G56110.1 68 GQFSEAEEHIIVKFHSVLGNR-WSLIAAQLP-GRTDNDVKNYWNT 110
789******************.*********.***********97 PP
|
| Publications
? help Back to Top |
- Li SF,Higginson T,Parish RW
A novel MYB-related gene from Arabidopsis thaliana expressed in developing anthers.
Plant Cell Physiol., 1999. 40(3): p. 343-7
[PMID:10353220] - Riechmann JL, et al.
Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes.
Science, 2000. 290(5499): p. 2105-10
[PMID:11118137] - Stracke R,Werber M,Weisshaar B
The R2R3-MYB gene family in Arabidopsis thaliana.
Curr. Opin. Plant Biol., 2001. 4(5): p. 447-56
[PMID:11597504] - Hanano S, et al.
Analysis of gene expression in Arabidopsis thaliana by array hybridization with genomic DNA fragments aligned along chromosomal regions.
Plant J., 2002. 30(2): p. 247-55
[PMID:12000460] - Higginson T,Li SF,Parish RW
AtMYB103 regulates tapetum and trichome development in Arabidopsis thaliana.
Plant J., 2003. 35(2): p. 177-92
[PMID:12848824] - Adham AR,Zolman BK,Millius A,Bartel B
Mutations in Arabidopsis acyl-CoA oxidase genes reveal distinct and overlapping roles in beta-oxidation.
Plant J., 2005. 41(6): p. 859-74
[PMID:15743450] - Cork JM,Purugganan MD
High-diversity genes in the Arabidopsis genome.
Genetics, 2005. 170(4): p. 1897-911
[PMID:15911589] - Yanhui C, et al.
The MYB transcription factor superfamily of Arabidopsis: expression analysis and phylogenetic comparison with the rice MYB family.
Plant Mol. Biol., 2006. 60(1): p. 107-24
[PMID:16463103] - Zhang W, et al.
Regulation of Arabidopsis tapetum development and function by DYSFUNCTIONAL TAPETUM1 (DYT1) encoding a putative bHLH transcription factor.
Development, 2006. 133(16): p. 3085-95
[PMID:16831835] - Li SF,Iacuone S,Parish RW
Suppression and restoration of male fertility using a transcription factor.
Plant Biotechnol. J., 2007. 5(2): p. 297-312
[PMID:17309685] - Zhang ZB, et al.
Transcription factor AtMYB103 is required for anther development by regulating tapetum development, callose dissolution and exine formation in Arabidopsis.
Plant J., 2007. 52(3): p. 528-38
[PMID:17727613] - Alves-Ferreira M, et al.
Global expression profiling applied to the analysis of Arabidopsis stamen development.
Plant Physiol., 2007. 145(3): p. 747-62
[PMID:17905860] - Zhu J, et al.
Defective in Tapetal development and function 1 is essential for anther development and tapetal function for microspore maturation in Arabidopsis.
Plant J., 2008. 55(2): p. 266-77
[PMID:18397379] - Zhong R,Lee C,Zhou J,McCarthy RL,Ye ZH
A battery of transcription factors involved in the regulation of secondary cell wall biosynthesis in Arabidopsis.
Plant Cell, 2008. 20(10): p. 2763-82
[PMID:18952777] - Zhu J, et al.
AtMYB103 is a crucial regulator of several pathways affecting Arabidopsis anther development.
Sci China Life Sci, 2010. 53(9): p. 1112-22
[PMID:21104372] - Phan HA,Iacuone S,Li SF,Parish RW
The MYB80 transcription factor is required for pollen development and the regulation of tapetal programmed cell death in Arabidopsis thaliana.
Plant Cell, 2011. 23(6): p. 2209-24
[PMID:21673079] - Zhu J,Lou Y,Xu X,Yang ZN
A genetic pathway for tapetum development and function in Arabidopsis.
J Integr Plant Biol, 2011. 53(11): p. 892-900
[PMID:21957980] - Phan HA,Li SF,Parish RW
MYB80, a regulator of tapetal and pollen development, is functionally conserved in crops.
Plant Mol. Biol., 2012. 78(1-2): p. 171-83
[PMID:22086333]
MYB103 is required for FERULATE-5-HYDROXYLASE expression and syringyl lignin biosynthesis in Arabidopsis stems.
Plant J., 2013. 73(1): p. 63-76
[PMID:22967312]- Xu Y,Iacuone S,Li SF,Parish RW
MYB80 homologues in Arabidopsis, cotton and Brassica: regulation and functional conservation in tapetal and pollen development.
BMC Plant Biol., 2014. 14: p. 278
[PMID:25311582] - Qian H, et al.
Trace concentrations of imazethapyr (IM) affect floral organs development and reproduction in Arabidopsis thaliana: IM-induced inhibition of key genes regulating anther and pollen biosynthesis.
Ecotoxicology, 2015. 24(1): p. 163-71
[PMID:25348600] - Jin J, et al.
An Arabidopsis Transcriptional Regulatory Map Reveals Distinct Functional and Evolutionary Features of Novel Transcription Factors.
Mol. Biol. Evol., 2015. 32(7): p. 1767-73
[PMID:25750178] - Xiong SX, et al.
The transcription factors MS188 and AMS form a complex to activate the expression of CYP703A2 for sporopollenin biosynthesis in Arabidopsis thaliana.
Plant J., 2016. 88(6): p. 936-946
[PMID:27460657] - Verma N,Burma PK
Regulation of tapetum-specific A9 promoter by transcription factors AtMYB80, AtMYB1 and AtMYB4 in Arabidopsis thaliana and Nicotiana tabacum.
Plant J., 2017. 92(3): p. 481-494
[PMID:28849604] - Li DD,Xue JS,Zhu J,Yang ZN
Gene Regulatory Network for Tapetum Development in Arabidopsis thaliana.
Front Plant Sci, 2017. 8: p. 1559
[PMID:28955355]
|





