 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT5G48560.1 | ||||||||
| Common Name | BHLH78, CIB2, EN86, K15N18.2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 498aa MW: 54877.8 Da PI: 6.3054 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 26 | 1.6e-08 | 311 | 358 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
+h+ +Er RR++i +++ L++l+P + +k Ka +L + ++Y++sLq
AT5G48560.1 311 SHSLAERVRREKIGERMKLLQDLVPGC----NKVTGKALMLDEIINYVQSLQ 358
8*************************9....677*****************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00083 | 4.37E-9 | 305 | 362 | No hit | No description |
| SuperFamily | SSF47459 | 3.14E-16 | 305 | 370 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 6.4E-16 | 307 | 372 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 15.37 | 307 | 357 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 1.1E-5 | 311 | 358 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 6.7E-10 | 313 | 363 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009637 | Biological Process | response to blue light | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000013 | anatomy | cauline leaf | ||||
| PO:0000230 | anatomy | inflorescence meristem | ||||
| PO:0000293 | anatomy | guard cell | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009025 | anatomy | vascular leaf | ||||
| PO:0009029 | anatomy | stamen | ||||
| PO:0009030 | anatomy | carpel | ||||
| PO:0009031 | anatomy | sepal | ||||
| PO:0009032 | anatomy | petal | ||||
| PO:0009046 | anatomy | flower | ||||
| PO:0009047 | anatomy | stem | ||||
| PO:0020100 | anatomy | hypocotyl | ||||
| PO:0020137 | anatomy | leaf apex | ||||
| PO:0025022 | anatomy | collective leaf structure | ||||
| PO:0025281 | anatomy | pollen | ||||
| PO:0007095 | developmental stage | LP.08 eight leaves visible stage | ||||
| PO:0007115 | developmental stage | LP.04 four leaves visible stage | ||||
| PO:0007123 | developmental stage | LP.06 six leaves visible stage | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| PO:0007616 | developmental stage | flowering stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 498 aa Download sequence Send to blast |
MDNELFMNTE FPPPPEMATH FEHQQSSSSA MMLNWALMDP NPHQDSSFLW EKSTEQQQQQ 60 SIFDSALSSL VSSPTPSNSN FSGGGGDGFL IRELIGKLGN IGNNNNNSGE IYGTPMSRSA 120 SCYATPMSSP PPPTNSNSQM MMNRTTPLTE FSADPGFAER AARFSCFGSR SFNGRTNTNL 180 PINNGNNMVN NSGKLTRVSS TPALKALVSP EVTPGGEFSR KRKSVPKGKS KENPISTASP 240 SPSFSKTAEK NGGKGGSKSS EEKGGKRRRE EEDDEEEEGE GEGNKSNNTK PPEPPKDYIH 300 VRARRGQATD SHSLAERVRR EKIGERMKLL QDLVPGCNKV TGKALMLDEI INYVQSLQRQ 360 VEFLSMKLSS VNDTRLDFNV DALVSKDVMI PSSNNRLHEE GLQSKSSSHH HQQQLNIYNN 420 NSQLLPNISS NNMMLQSPMN SLETSTLARS FTHLPTLTQF TDSISQYQMF SEEDLQSIVG 480 MGVAENPNNE SQHMKIEL |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 218 | 223 | SRKRKS |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| At.43769 | 0.0 | root| silique | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 30695486 | 0.0 | ||||
| Genevisible | 248708_at | 0.0 | ||||
| Expression Atlas | AT5G48560 | - | ||||
| AtGenExpress | AT5G48560 | - | ||||
| ATTED-II | AT5G48560 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed constitutively in roots, leaves, stems, and flowers. {ECO:0000269|PubMed:12679534}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that binds DNA to G box 5'-CACGTG-3' and to E-box 5'-CANNTG-3' (By similarity). Binds to chromatin DNA of the FT gene and promotes its expression, and thus triggers flowering in response to blue light (PubMed:24130508). {ECO:0000250|UniProtKB:Q8GY61, ECO:0000269|PubMed:24130508}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00036 | PBM | 25215497 | Download |
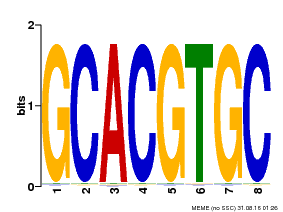
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT5G48560.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Interaction ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Intact With | |||||
| IntAct | Search Q9FJL4 | |||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT5G48560 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT002945 | 0.0 | BT002945.1 Arabidopsis thaliana clone RAFL14-96-A17 (R20245) unknown protein (At5g48560) mRNA, complete cds. | |||
| GenBank | BT005637 | 0.0 | BT005637.1 Arabidopsis thaliana clone U20245 unknown protein (At5g48560) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_199667.1 | 0.0 | basic helix-loop-helix (bHLH) DNA-binding superfamily protein | ||||
| Swissprot | Q9FJL4 | 0.0 | BH078_ARATH; Transcription factor bHLH78 | ||||
| TrEMBL | A0A178UME9 | 0.0 | A0A178UME9_ARATH; Uncharacterized protein | ||||
| STRING | AT5G48560.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2332 | 27 | 75 | Representative plant | OGRP58 | 16 | 313 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT5G48560.1 |
| Entrez Gene | 834912 |
| iHOP | AT5G48560 |
| wikigenes | AT5G48560 |



