 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT5G05090.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 266aa MW: 29441.1 Da PI: 8.0803 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 97 | 1.4e-30 | 81 | 134 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
+prl+Wtp+LH+rFv+av++L G+++A+Pkti++lm+v+gLt+e+v+SHLQkYRl
AT5G05090.1 81 RPRLVWTPQLHKRFVDAVAHL-GIKNAVPKTIMQLMSVDGLTRENVASHLQKYRL 134
59*******************.********************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 11.059 | 78 | 137 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.33E-20 | 78 | 138 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 8.4E-31 | 80 | 139 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.6E-24 | 82 | 134 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 9.4E-8 | 84 | 133 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000013 | anatomy | cauline leaf | ||||
| PO:0000037 | anatomy | shoot apex | ||||
| PO:0000230 | anatomy | inflorescence meristem | ||||
| PO:0000293 | anatomy | guard cell | ||||
| PO:0008019 | anatomy | leaf lamina base | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009009 | anatomy | plant embryo | ||||
| PO:0009025 | anatomy | vascular leaf | ||||
| PO:0009029 | anatomy | stamen | ||||
| PO:0009030 | anatomy | carpel | ||||
| PO:0009031 | anatomy | sepal | ||||
| PO:0009032 | anatomy | petal | ||||
| PO:0009046 | anatomy | flower | ||||
| PO:0009047 | anatomy | stem | ||||
| PO:0009052 | anatomy | flower pedicel | ||||
| PO:0020030 | anatomy | cotyledon | ||||
| PO:0020038 | anatomy | petiole | ||||
| PO:0020100 | anatomy | hypocotyl | ||||
| PO:0020137 | anatomy | leaf apex | ||||
| PO:0025022 | anatomy | collective leaf structure | ||||
| PO:0025195 | anatomy | pollen tube cell | ||||
| PO:0025281 | anatomy | pollen | ||||
| PO:0001054 | developmental stage | vascular leaf senescent stage | ||||
| PO:0001185 | developmental stage | plant embryo globular stage | ||||
| PO:0004507 | developmental stage | plant embryo bilateral stage | ||||
| PO:0007064 | developmental stage | LP.12 twelve leaves visible stage | ||||
| PO:0007095 | developmental stage | LP.08 eight leaves visible stage | ||||
| PO:0007098 | developmental stage | LP.02 two leaves visible stage | ||||
| PO:0007103 | developmental stage | LP.10 ten leaves visible stage | ||||
| PO:0007115 | developmental stage | LP.04 four leaves visible stage | ||||
| PO:0007123 | developmental stage | LP.06 six leaves visible stage | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| PO:0007616 | developmental stage | flowering stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 266 aa Download sequence Send to blast |
MREEDSNWFA KWEEELPSPE ELIPLSQSLI TPDLAIAFDL HRNNNSNSGQ PLPQTTPPQP 60 NSSAEIAGDS TGDEPARTLK RPRLVWTPQL HKRFVDAVAH LGIKNAVPKT IMQLMSVDGL 120 TRENVASHLQ KYRLYLKRMK SGGGGGGSGD SDHLFASSPV PPHFLHPTSR QSSDLFIPSF 180 VPISTQQQHI AAPPSQFLHR QISAVNFTSP TKATDQAMFL ARQQSELQQP VFKPSSLHLH 240 SQVANYTQDL KSGAKTVLTL FPTRDD |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5lxu_A | 4e-32 | 81 | 137 | 1 | 57 | Transcription factor LUX |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| At.33034 | 0.0 | flower | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 30680790 | 0.0 | ||||
| Genevisible | 250810_at | 0.0 | ||||
| Expression Atlas | AT5G05090 | - | ||||
| AtGenExpress | AT5G05090 | - | ||||
| ATTED-II | AT5G05090 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, leaves, stems, petioles, filaments, stigma, pedicels, sepals, anthers, petals, and siliques. {ECO:0000269|PubMed:20331973}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that acts as a negative regulator of freezing tolerance via a CBF-independent pathway. {ECO:0000269|PubMed:20331973}. | |||||
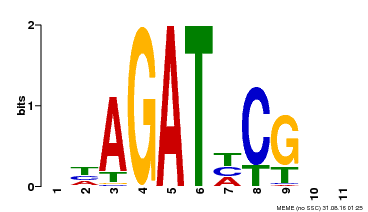
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00056 | PBM | 25215497 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT5G05090.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by drought, salt and abscisic acid (ABA). Down-regulated by cold. {ECO:0000269|PubMed:20331973}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT5G05090 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB005245 | 0.0 | AB005245.2 Arabidopsis thaliana genomic DNA, chromosome 5, P1 clone:MUG13. | |||
| GenBank | AY093188 | 0.0 | AY093188.1 Arabidopsis thaliana unknown protein (At5g05090) mRNA, complete cds. | |||
| GenBank | BT006289 | 0.0 | BT006289.1 Arabidopsis thaliana At5g05090 mRNA, complete cds. | |||
| GenBank | CP002688 | 0.0 | CP002688.1 Arabidopsis thaliana chromosome 5 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_196128.1 | 0.0 | Homeodomain-like superfamily protein | ||||
| Swissprot | O22210 | 6e-55 | MYBC1_ARATH; Transcription factor MYBC1 | ||||
| TrEMBL | Q9FF65 | 0.0 | Q9FF65_ARATH; At5g05090 | ||||
| STRING | AT5G05090.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1549 | 27 | 91 | Representative plant | OGRP1157 | 17 | 53 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT5G05090.1 |
| Entrez Gene | 830391 |
| iHOP | AT5G05090 |
| wikigenes | AT5G05090 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



