| Signature Domain? help Back to Top |
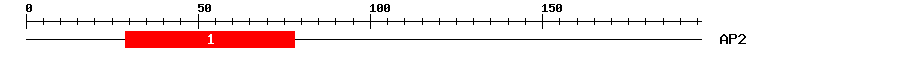 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | AP2 | 58.8 | 1.3e-18 | 29 | 78 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
ykG+r +k +g+WvAeIr+p +kr r++lg++ t+e Aa+a++ a +l+g
AT4G36900.1 29 PYKGIRMRK-WGKWVAEIREP---NKRSRIWLGSYSTPEAAARAYDTAVFYLRG 78
69*****99.**********9...336*************************98 PP
|
| Publications
? help Back to Top |
- Terryn N, et al.
Evidence for an ancient chromosomal duplication in Arabidopsis thaliana by sequencing and analyzing a 400-kb contig at the APETALA2 locus on chromosome 4.
FEBS Lett., 1999. 445(2-3): p. 237-45
[PMID:10094464] - Riechmann JL, et al.
Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes.
Science, 2000. 290(5499): p. 2105-10
[PMID:11118137] - Haake V, et al.
Transcription factor CBF4 is a regulator of drought adaptation in Arabidopsis.
Plant Physiol., 2002. 130(2): p. 639-48
[PMID:12376631] - Brown RL,Kazan K,McGrath KC,Maclean DJ,Manners JM
A role for the GCC-box in jasmonate-mediated activation of the PDF1.2 gene of Arabidopsis.
Plant Physiol., 2003. 132(2): p. 1020-32
[PMID:12805630] - Dal Bosco C, et al.
Inactivation of the chloroplast ATP synthase gamma subunit results in high non-photochemical fluorescence quenching and altered nuclear gene expression in Arabidopsis thaliana.
J. Biol. Chem., 2004. 279(2): p. 1060-9
[PMID:14576160] - Yamada K, et al.
Empirical analysis of transcriptional activity in the Arabidopsis genome.
Science, 2003. 302(5646): p. 842-6
[PMID:14593172] - Jiao Y, et al.
A genome-wide analysis of blue-light regulation of Arabidopsis transcription factor gene expression during seedling development.
Plant Physiol., 2003. 133(4): p. 1480-93
[PMID:14605227] - Johnson MA, et al.
Arabidopsis hapless mutations define essential gametophytic functions.
Genetics, 2004. 168(2): p. 971-82
[PMID:15514068] - Lee BH,Henderson DA,Zhu JK
The Arabidopsis cold-responsive transcriptome and its regulation by ICE1.
Plant Cell, 2005. 17(11): p. 3155-75
[PMID:16214899] - Duarte JM, et al.
Expression pattern shifts following duplication indicative of subfunctionalization and neofunctionalization in regulatory genes of Arabidopsis.
Mol. Biol. Evol., 2006. 23(2): p. 469-78
[PMID:16280546] - Nakano T,Suzuki K,Fujimura T,Shinshi H
Genome-wide analysis of the ERF gene family in Arabidopsis and rice.
Plant Physiol., 2006. 140(2): p. 411-32
[PMID:16407444] - Truman W,de Zabala MT,Grant M
Type III effectors orchestrate a complex interplay between transcriptional networks to modify basal defence responses during pathogenesis and resistance.
Plant J., 2006. 46(1): p. 14-33
[PMID:16553893] - Xiong Y,Fei SZ
Functional and phylogenetic analysis of a DREB/CBF-like gene in perennial ryegrass (Lolium perenne L.).
Planta, 2006. 224(4): p. 878-88
[PMID:16614820] - Rossel JB, et al.
Systemic and intracellular responses to photooxidative stress in Arabidopsis.
Plant Cell, 2007. 19(12): p. 4091-110
[PMID:18156220] - Welling A,Palva ET
Involvement of CBF transcription factors in winter hardiness in birch.
Plant Physiol., 2008. 147(3): p. 1199-211
[PMID:18467468] - Tsutsui T, et al.
DEAR1, a transcriptional repressor of DREB protein that mediates plant defense and freezing stress responses in Arabidopsis.
J. Plant Res., 2009. 122(6): p. 633-43
[PMID:19618250] - Ding Y, et al.
Four distinct types of dehydration stress memory genes in Arabidopsis thaliana.
BMC Plant Biol., 2013. 13: p. 229
[PMID:24377444] - Coego A, et al.
The TRANSPLANTA collection of Arabidopsis lines: a resource for functional analysis of transcription factors based on their conditional overexpression.
Plant J., 2014. 77(6): p. 944-53
[PMID:24456507] - Okamuro JK,Caster B,Villarroel R,Van Montagu M,Jofuku KD
The AP2 domain of APETALA2 defines a large new family of DNA binding proteins in Arabidopsis.
Proc. Natl. Acad. Sci. U.S.A., 1997. 94(13): p. 7076-81
[PMID:9192694]
|





