 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT4G36780.1 | ||||||||
| Common Name | BEH2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | BES1 | ||||||||
| Protein Properties | Length: 265aa MW: 28775.8 Da PI: 11.3329 | ||||||||
| Description | BES1/BZR1 homolog 2 | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | DUF822 | 220.3 | 4e-68 | 13 | 153 | 2 | 145 |
DUF822 2 gsgrkptwkErEnnkrRERrRRaiaakiyaGLRaqGnyklpkraDnneVlkALcreAGwvvedDGttyrkgskpleeaeaagssasaspesslqsslk 99
+sgr+ptwkErEnnk+RERrRRai+akiy+GLRaqGnyklpk++DnneVlkALc eAGw+vedDGttyrkg+kp+ ++++ g+ ++ s++ss+q s++
AT4G36780.1 13 SSGRTPTWKERENNKKRERRRRAITAKIYSGLRAQGNYKLPKHCDNNEVLKALCLEAGWIVEDDGTTYRKGFKPP-ASDISGTPTNFSTNSSIQPSPQ 109
89*************************************************************************.********************** PP
DUF822 100 ssalaspvesysaspksssfpspssldsislasaasllpvlsvlsl 145
ssa++sp++sy+ sp sssfpsps++d ++++ llp+l++ ++
AT4G36780.1 110 SSAFPSPAPSYHGSPVSSSFPSPSRYDGNPSS--YLLLPFLHNIAS 153
*****************************987..899999998765 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF05687 | 2.5E-62 | 13 | 137 | IPR008540 | BES1/BZR1 plant transcription factor, N-terminal |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000013 | anatomy | cauline leaf | ||||
| PO:0000037 | anatomy | shoot apex | ||||
| PO:0000230 | anatomy | inflorescence meristem | ||||
| PO:0000293 | anatomy | guard cell | ||||
| PO:0008019 | anatomy | leaf lamina base | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009006 | anatomy | shoot system | ||||
| PO:0009009 | anatomy | plant embryo | ||||
| PO:0009010 | anatomy | seed | ||||
| PO:0009025 | anatomy | vascular leaf | ||||
| PO:0009029 | anatomy | stamen | ||||
| PO:0009030 | anatomy | carpel | ||||
| PO:0009031 | anatomy | sepal | ||||
| PO:0009032 | anatomy | petal | ||||
| PO:0009046 | anatomy | flower | ||||
| PO:0009047 | anatomy | stem | ||||
| PO:0009052 | anatomy | flower pedicel | ||||
| PO:0020030 | anatomy | cotyledon | ||||
| PO:0020038 | anatomy | petiole | ||||
| PO:0020100 | anatomy | hypocotyl | ||||
| PO:0020137 | anatomy | leaf apex | ||||
| PO:0025022 | anatomy | collective leaf structure | ||||
| PO:0025281 | anatomy | pollen | ||||
| PO:0001054 | developmental stage | vascular leaf senescent stage | ||||
| PO:0001078 | developmental stage | plant embryo cotyledonary stage | ||||
| PO:0001081 | developmental stage | mature plant embryo stage | ||||
| PO:0001185 | developmental stage | plant embryo globular stage | ||||
| PO:0004507 | developmental stage | plant embryo bilateral stage | ||||
| PO:0007064 | developmental stage | LP.12 twelve leaves visible stage | ||||
| PO:0007095 | developmental stage | LP.08 eight leaves visible stage | ||||
| PO:0007098 | developmental stage | LP.02 two leaves visible stage | ||||
| PO:0007103 | developmental stage | LP.10 ten leaves visible stage | ||||
| PO:0007115 | developmental stage | LP.04 four leaves visible stage | ||||
| PO:0007123 | developmental stage | LP.06 six leaves visible stage | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| PO:0007616 | developmental stage | flowering stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 265 aa Download sequence Send to blast |
MAAGGGGGGG GSSSGRTPTW KERENNKKRE RRRRAITAKI YSGLRAQGNY KLPKHCDNNE 60 VLKALCLEAG WIVEDDGTTY RKGFKPPASD ISGTPTNFST NSSIQPSPQS SAFPSPAPSY 120 HGSPVSSSFP SPSRYDGNPS SYLLLPFLHN IASSIPANLP PLRISNSAPV TPPLSSPTSR 180 GSKRKLTSEQ LPNGGSLHVL RHPLFAISAP SRLVVLVTKR HLQYRNVMSP RRIRSRIQEG 240 GSISNLLLLL HQHLTLFSKL LWPLI |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5zd4_A | 2e-27 | 16 | 94 | 372 | 450 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| 5zd4_B | 2e-27 | 16 | 94 | 372 | 450 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| 5zd4_C | 2e-27 | 16 | 94 | 372 | 450 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| 5zd4_D | 2e-27 | 16 | 94 | 372 | 450 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| At.22285 | 0.0 | root | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 240256203 | 0.0 | ||||
| Genevisible | 246284_at | 0.0 | ||||
| Expression Atlas | AT4G36780 | - | ||||
| AtGenExpress | AT4G36780 | - | ||||
| ATTED-II | AT4G36780 | - | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
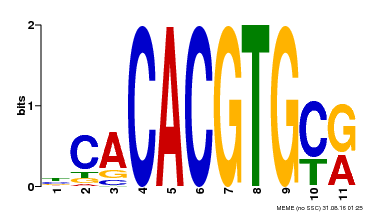
| Motif ID | Method | Source | Motif file |
| MP00474 | DAP | 27203113 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT4G36780.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Regulation -- Hormone ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hormone | |||||
| AHD | brassinosteroid | |||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT4G36780 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY050394 | 0.0 | AY050394.1 Arabidopsis thaliana AT4g36780/C7A10_580 mRNA, complete cds. | |||
| GenBank | AY097357 | 0.0 | AY097357.1 Arabidopsis thaliana AT4g36780/C7A10_580 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_195396.4 | 0.0 | BES1/BZR1 homolog 2 | ||||
| Swissprot | Q94A43 | 1e-150 | BEH2_ARATH; BES1/BZR1 homolog protein 2 | ||||
| TrEMBL | A0A178V2S6 | 1e-146 | A0A178V2S6_ARATH; BEH2 | ||||
| STRING | AT4G36780.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM11512 | 23 | 33 | Representative plant | OGRP3719 | 10 | 24 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT4G36780.1 |
| Entrez Gene | 829831 |
| iHOP | AT4G36780 |
| wikigenes | AT4G36780 |



