| Signature Domain? help Back to Top |
 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | SRF-TF | 92.3 | 2.3e-29 | 9 | 59 | 1 | 51 |
S---SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-TTSEEEEEE- CS
SRF-TF 1 krienksnrqvtfskRrngilKKAeELSvLCdaevaviifsstgklyeyss 51
krien + rqvtfskRrng+lKKA+ELSvLCdaeva++ifs++ klye+ss
AT4G22950.1 9 KRIENATSRQVTFSKRRNGLLKKAFELSVLCDAEVALVIFSPRSKLYEFSS 59
79***********************************************96 PP
|
| 2 | K-box | 79.4 | 8.8e-27 | 79 | 170 | 7 | 98 |
K-box 7 ksleeakaeslqqelakLkkeienLqreqRhllGedLesLslkeLqqLeqqLekslkkiRskKnellleqieelqkkekelqeenkaLrkkl 98
+++ + ++++ + e + L k+ie+L+ ++R+llGe+++ +s++eLqqLe+qL++sl++iR+kK++ll+e+ie+l+ e++l +enk+L++k+
AT4G22950.1 79 NHKRNDNSQQARDETSGLTKKIEQLEISKRKLLGEGIDACSIEELQQLENQLDRSLSRIRAKKYQLLREEIEKLKAEERNLVKENKDLKEKW 170
4467788999*******************************************************************************996 PP
|
| Protein Features
? help Back to Top |
 |
| Database |
Entry ID |
E-value |
Start |
End |
InterPro ID |
Description |
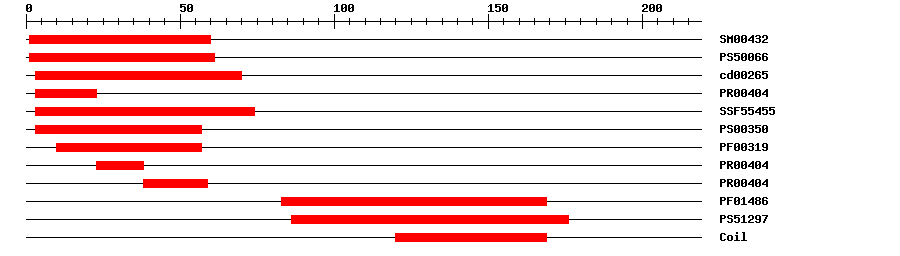
| SMART | SM00432 | 3.5E-39 | 1 | 60 | IPR002100 | Transcription factor, MADS-box |
| PROSITE profile | PS50066 | 31.286 | 1 | 61 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00265 | 1.85E-39 | 3 | 70 | No hit | No description |
| PRINTS | PR00404 | 4.4E-30 | 3 | 23 | IPR002100 | Transcription factor, MADS-box |
| SuperFamily | SSF55455 | 1.44E-31 | 3 | 74 | IPR002100 | Transcription factor, MADS-box |
| PROSITE pattern | PS00350 | 0 | 3 | 57 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF00319 | 4.8E-26 | 10 | 57 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 4.4E-30 | 23 | 38 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 4.4E-30 | 38 | 59 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF01486 | 3.7E-26 | 83 | 169 | IPR002487 | Transcription factor, K-box |
| PROSITE profile | PS51297 | 14.845 | 86 | 176 | IPR002487 | Transcription factor, K-box |
| Publications
? help Back to Top |
- Alvarez-Buylla ER, et al.
MADS-box gene evolution beyond flowers: expression in pollen, endosperm, guard cells, roots and trichomes.
Plant J., 2000. 24(4): p. 457-66
[PMID:11115127] - Riechmann JL, et al.
Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes.
Science, 2000. 290(5499): p. 2105-10
[PMID:11118137] - Parenicová L, et al.
Molecular and phylogenetic analyses of the complete MADS-box transcription factor family in Arabidopsis: new openings to the MADS world.
Plant Cell, 2003. 15(7): p. 1538-51
[PMID:12837945] - Kofuji R, et al.
Evolution and divergence of the MADS-box gene family based on genome-wide expression analyses.
Mol. Biol. Evol., 2003. 20(12): p. 1963-77
[PMID:12949148] - Yamada K, et al.
Empirical analysis of transcriptional activity in the Arabidopsis genome.
Science, 2003. 302(5646): p. 842-6
[PMID:14593172] - Czechowski T,Bari RP,Stitt M,Scheible WR,Udvardi MK
Real-time RT-PCR profiling of over 1400 Arabidopsis transcription factors: unprecedented sensitivity reveals novel root- and shoot-specific genes.
Plant J., 2004. 38(2): p. 366-79
[PMID:15078338] - De Paepe A,Vuylsteke M,Van Hummelen P,Zabeau M,Van Der Straeten D
Transcriptional profiling by cDNA-AFLP and microarray analysis reveals novel insights into the early response to ethylene in Arabidopsis.
Plant J., 2004. 39(4): p. 537-59
[PMID:15272873] - de Folter S, et al.
Comprehensive interaction map of the Arabidopsis MADS Box transcription factors.
Plant Cell, 2005. 17(5): p. 1424-33
[PMID:15805477] - Pina C,Pinto F,Feij
Gene family analysis of the Arabidopsis pollen transcriptome reveals biological implications for cell growth, division control, and gene expression regulation.
Plant Physiol., 2005. 138(2): p. 744-56
[PMID:15908605] - Gan Y,Filleur S,Rahman A,Gotensparre S,Forde BG
Nutritional regulation of ANR1 and other root-expressed MADS-box genes in Arabidopsis thaliana.
Planta, 2005. 222(4): p. 730-42
[PMID:16021502] - Duarte JM, et al.
Expression pattern shifts following duplication indicative of subfunctionalization and neofunctionalization in regulatory genes of Arabidopsis.
Mol. Biol. Evol., 2006. 23(2): p. 469-78
[PMID:16280546] - Schönrock N, et al.
Polycomb-group proteins repress the floral activator AGL19 in the FLC-independent vernalization pathway.
Genes Dev., 2006. 20(12): p. 1667-78
[PMID:16778081] - Pien S, et al.
ARABIDOPSIS TRITHORAX1 dynamically regulates FLOWERING LOCUS C activation via histone 3 lysine 4 trimethylation.
Plant Cell, 2008. 20(3): p. 580-8
[PMID:18375656] - Alexandre CM,Hennig L
FLC or not FLC: the other side of vernalization.
J. Exp. Bot., 2008. 59(6): p. 1127-35
[PMID:18390846] - Liang S, et al.
Transcriptional Regulations on the Low-Temperature-Induced Floral Transition in an Orchidaceae Species, Dendrobium nobile: An Expressed Sequence Tags Analysis.
Comp. Funct. Genomics, 2012. 2012: p. 757801
[PMID:22550428] - Kim W,Latrasse D,Servet C,Zhou DX
Arabidopsis histone deacetylase HDA9 regulates flowering time through repression of AGL19.
Biochem. Biophys. Res. Commun., 2013. 432(2): p. 394-8
[PMID:23237803] - Suter L,Rüegg M,Zemp N,Hennig L,Widmer A
Gene regulatory variation mediates flowering responses to vernalization along an altitudinal gradient in Arabidopsis.
Plant Physiol., 2014. 166(4): p. 1928-42
[PMID:25339407] - Kang MJ,Jin HS,Noh YS,Noh B
Repression of flowering under a noninductive photoperiod by the HDA9-AGL19-FT module in Arabidopsis.
New Phytol., 2015. 206(1): p. 281-94
[PMID:25406502]
|




