 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT4G00250.1 | ||||||||
| Common Name | A_IG005I10.12, F5I10.12 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | GeBP | ||||||||
| Protein Properties | Length: 319aa MW: 35513.3 Da PI: 9.4264 | ||||||||
| Description | DNA-binding storekeeper protein-related transcriptional regulator | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | DUF573 | 108.1 | 6.4e-34 | 114 | 204 | 2 | 98 |
DUF573 2 lfqrlwseeDeivlLqGlidfkaktgkspsddidafyefvkksisfkvsksqlveKirrLKkKfkkkvkkaksgkepsfskehdqkifelskkiWgs 98
++qr+wseeDei+lLq++idfka+tg+sp+d+ +af++++kksisf+vs+ q+++KirrLK+K+ ++k+ +s +++hd+k+++l+ iWgs
AT4G00250.1 114 NPQRVWSEEDEISLLQAVIDFKAETGTSPWDHKNAFFDIAKKSISFDVSHVQFFDKIRRLKNKYFVNRKN------KSGESNHDKKCLGLAVLIWGS 204
68******************************************************************99......667789*************95 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF04504 | 1.0E-24 | 116 | 203 | IPR007592 | Protein of unknown function DUF573 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010629 | Biological Process | negative regulation of gene expression | ||||
| GO:0071333 | Biological Process | cellular response to glucose stimulus | ||||
| GO:0005623 | Cellular Component | cell | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 319 aa Download sequence Send to blast |
MAPLESPATA SSSEVESSSE EIFKSSSEES KPKDPVTVPS SKTLKSPSAA VNSKTDSSDD 60 SEKQSFVLTR RKKKEGAAES PAVKSGKKRA GEGSTSRDMH VKRVKKEDDN KKANPQRVWS 120 EEDEISLLQA VIDFKAETGT SPWDHKNAFF DIAKKSISFD VSHVQFFDKI RRLKNKYFVN 180 RKNKSGESNH DKKCLGLAVL IWGSDGMNVE SPVKKDESIL VKGKANSKEK KVEKPLVIED 240 EQVILGADSE WFEESFLVPV IANLGLDEYS VKKKWSKVSL ETKKKIQEKM KVVDAKKCEL 300 LLAEMDVLKD VTSVLAQTN |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 69 | 74 | RRKKKE |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| At.34582 | 0.0 | seed | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| Genevisible | 255668_s_at | 1e-161 | ||||
| Expression Atlas | AT4G00250 | - | ||||
| AtGenExpress | AT4G00250 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed strongly in leaves and flowers, weakly in roots, and very weakly in stems. {ECO:0000269|PubMed:27031427}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription repressor that binds DNA in a sequence-specific manner, 5'-GCCT-3', to regulate the expression of PGR. Acts as a modulatory component for the glucose-triggered developmental leaf growth process. {ECO:0000269|PubMed:27031427}. | |||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
|
||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00420 | DAP | 27203113 | Download |
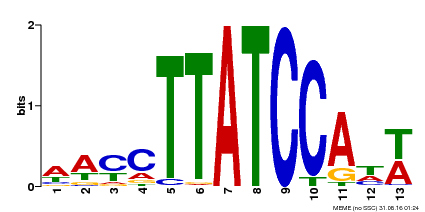
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT4G00250.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated upon glucose treatment. {ECO:0000269|PubMed:27031427}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT4G00250 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AL161471 | 0.0 | AL161471.2 Arabidopsis thaliana DNA chromosome 4, contig fragment No. 1. | |||
| GenBank | BT003961 | 0.0 | BT003961.1 Arabidopsis thaliana clone RAFL15-21-E22 (R20787) unknown protein (At4g00250) mRNA, complete cds. | |||
| GenBank | BT005666 | 0.0 | BT005666.1 Arabidopsis thaliana clone U20787 unknown protein (At4g00250) mRNA, complete cds. | |||
| GenBank | CP002687 | 0.0 | CP002687.1 Arabidopsis thaliana chromosome 4 sequence. | |||
| GenBank | F5I10 | 0.0 | AF195115.1 Arabidopsis thaliana BAC F5I10. | |||
| GenBank | IG005I10 | 0.0 | AF013293.1 Arabidopsis thaliana BAC IG005I10. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_567161.1 | 0.0 | DNA-binding storekeeper protein-related transcriptional regulator | ||||
| Swissprot | O23077 | 0.0 | STKLN_ARATH; Transcription factor STKL2 | ||||
| TrEMBL | A0A178UZP9 | 1e-151 | A0A178UZP9_ARATH; Uncharacterized protein | ||||
| STRING | AT4G00250.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM313 | 28 | 181 | Representative plant | OGRP4222 | 5 | 23 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT4G00250.1 |
| Entrez Gene | 828046 |
| iHOP | AT4G00250 |
| wikigenes | AT4G00250 |



