| Signature Domain? help Back to Top |
 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | SRF-TF | 86.3 | 1.8e-27 | 9 | 59 | 1 | 51 |
S---SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-TTSEEEEEE- CS
SRF-TF 1 krienksnrqvtfskRrngilKKAeELSvLCdaevaviifsstgklyeyss 51
k+ien rqvtfskRrng++KKA+ELS+LCdaeva+iifsstgk+y++ss
AT3G57390.1 9 KKIENINSRQVTFSKRRNGLIKKAKELSILCDAEVALIIFSSTGKIYDFSS 59
68***********************************************96 PP
|
| 2 | K-box | 54.7 | 4.3e-19 | 97 | 179 | 17 | 99 |
K-box 17 lqqelakLkkeienLqreqRhllGedLesLslkeLqqLeqqLekslkkiRskKnellleqieelqkkekelqeenkaLrkkle 99
++ + +k e+e Lq + +l+G++Le +s+ +L +Le+qL++sl +++++K+++ll+qie+++ +ek++ een+ Lrk++e
AT3G57390.1 97 VLRNDDSMKGELERLQLAIERLKGKELEGMSFPDLISLENQLNESLHSVKDQKTQILLNQIERSRIQEKKALEENQILRKQVE 179
55566899************************************************************************997 PP
|
| Protein Features
? help Back to Top |
 |
| Database |
Entry ID |
E-value |
Start |
End |
InterPro ID |
Description |
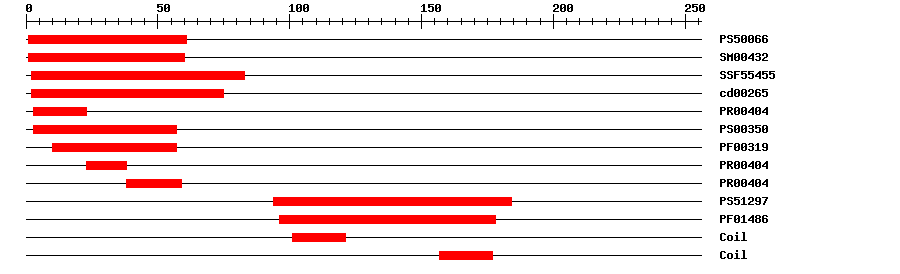
| PROSITE profile | PS50066 | 31.833 | 1 | 61 | IPR002100 | Transcription factor, MADS-box |
| SMART | SM00432 | 2.2E-39 | 1 | 60 | IPR002100 | Transcription factor, MADS-box |
| SuperFamily | SSF55455 | 4.58E-31 | 2 | 83 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00265 | 9.25E-41 | 2 | 75 | No hit | No description |
| PRINTS | PR00404 | 6.2E-29 | 3 | 23 | IPR002100 | Transcription factor, MADS-box |
| PROSITE pattern | PS00350 | 0 | 3 | 57 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF00319 | 1.1E-25 | 10 | 57 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 6.2E-29 | 23 | 38 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 6.2E-29 | 38 | 59 | IPR002100 | Transcription factor, MADS-box |
| PROSITE profile | PS51297 | 13.259 | 94 | 184 | IPR002487 | Transcription factor, K-box |
| Pfam | PF01486 | 2.2E-16 | 96 | 178 | IPR002487 | Transcription factor, K-box |
| Expression --
Description ? help
Back to Top |
| Source |
Description |
| Uniprot | DEVELOPMENTAL STAGE: During the reproductive phase, accumulates in immature buds and at the base of the floral organs, and in the receptacle, ovules, anther filaments, and stigma and style of open flowers. Later observed in sporogenous tissue of anthers. During male gametogenesis, expressed in the microspores before they separate from each other. Later present at high levels within pollen grains up to stage 13 of flower development, when anthers dehisce. During carpel development, first detected in developing ovules. After fertilization, confined to globular structures or nodules of proliferating free nuclear endosperm required for embryo development. Disappears from the endosperm at to the heart stage of embryo development, when very little nuclear endosperm remains. Never detected in developing embryos at any stage. In young seedlings, present everywhere except in a portion of the hypocotyl and in newly emerging leaves. {ECO:0000269|PubMed:11115127, ECO:0000269|PubMed:17521410}. |
| Uniprot | TISSUE SPECIFICITY: Mostly expressed in pollen, roots, flowers and siliques, and to a lower extent, in stems and leaves. Expressed in the endosperm and in developing male and female gametophytes. Also present in seedlings. {ECO:0000269|PubMed:11115127, ECO:0000269|PubMed:12837945, ECO:0000269|PubMed:12949148, ECO:0000269|PubMed:16028119, ECO:0000269|PubMed:17521410}. |
| Publications
? help Back to Top |
- Alvarez-Buylla ER, et al.
MADS-box gene evolution beyond flowers: expression in pollen, endosperm, guard cells, roots and trichomes.
Plant J., 2000. 24(4): p. 457-66
[PMID:11115127] - Riechmann JL, et al.
Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes.
Science, 2000. 290(5499): p. 2105-10
[PMID:11118137] - Ma L, et al.
Genomic evidence for COP1 as a repressor of light-regulated gene expression and development in Arabidopsis.
Plant Cell, 2002. 14(10): p. 2383-98
[PMID:12368493] - Zik M,Irish VF
Global identification of target genes regulated by APETALA3 and PISTILLATA floral homeotic gene action.
Plant Cell, 2003. 15(1): p. 207-22
[PMID:12509532] - Parenicová L, et al.
Molecular and phylogenetic analyses of the complete MADS-box transcription factor family in Arabidopsis: new openings to the MADS world.
Plant Cell, 2003. 15(7): p. 1538-51
[PMID:12837945] - Kofuji R, et al.
Evolution and divergence of the MADS-box gene family based on genome-wide expression analyses.
Mol. Biol. Evol., 2003. 20(12): p. 1963-77
[PMID:12949148] - Pina C,Pinto F,Feij
Gene family analysis of the Arabidopsis pollen transcriptome reveals biological implications for cell growth, division control, and gene expression regulation.
Plant Physiol., 2005. 138(2): p. 744-56
[PMID:15908605] - Gan Y,Filleur S,Rahman A,Gotensparre S,Forde BG
Nutritional regulation of ANR1 and other root-expressed MADS-box genes in Arabidopsis thaliana.
Planta, 2005. 222(4): p. 730-42
[PMID:16021502] - Lehti-Shiu MD,Adamczyk BJ,Fernandez DE
Expression of MADS-box genes during the embryonic phase in Arabidopsis.
Plant Mol. Biol., 2005. 58(1): p. 89-107
[PMID:16028119] - He XJ, et al.
AtNAC2, a transcription factor downstream of ethylene and auxin signaling pathways, is involved in salt stress response and lateral root development.
Plant J., 2005. 44(6): p. 903-16
[PMID:16359384] - Truman W,de Zabala MT,Grant M
Type III effectors orchestrate a complex interplay between transcriptional networks to modify basal defence responses during pathogenesis and resistance.
Plant J., 2006. 46(1): p. 14-33
[PMID:16553893] - Helliwell CA,Wood CC,Robertson M,James Peacock W,Dennis ES
The Arabidopsis FLC protein interacts directly in vivo with SOC1 and FT chromatin and is part of a high-molecular-weight protein complex.
Plant J., 2006. 46(2): p. 183-92
[PMID:16623882] - Kim J,Shiu SH,Thoma S,Li WH,Patterson SE
Patterns of expansion and expression divergence in the plant polygalacturonase gene family.
Genome Biol., 2006. 7(9): p. R87
[PMID:17010199] - Verelst W,Saedler H,Münster T
MIKC* MADS-protein complexes bind motifs enriched in the proximal region of late pollen-specific Arabidopsis promoters.
Plant Physiol., 2007. 143(1): p. 447-60
[PMID:17071640] - Adamczyk BJ,Lehti-Shiu MD,Fernandez DE
The MADS domain factors AGL15 and AGL18 act redundantly as repressors of the floral transition in Arabidopsis.
Plant J., 2007. 50(6): p. 1007-19
[PMID:17521410] - Hill K,Wang H,Perry SE
A transcriptional repression motif in the MADS factor AGL15 is involved in recruitment of histone deacetylase complex components.
Plant J., 2008. 53(1): p. 172-85
[PMID:17999645] - Verelst W, et al.
MADS-complexes regulate transcriptome dynamics during pollen maturation.
Genome Biol., 2007. 8(11): p. R249
[PMID:18034896] - Thakare D,Tang W,Hill K,Perry SE
The MADS-domain transcriptional regulator AGAMOUS-LIKE15 promotes somatic embryo development in Arabidopsis and soybean.
Plant Physiol., 2008. 146(4): p. 1663-72
[PMID:18305206] - Wang Y, et al.
Transcriptome analyses show changes in gene expression to accompany pollen germination and tube growth in Arabidopsis.
Plant Physiol., 2008. 148(3): p. 1201-11
[PMID:18775970] - Pouteau S, et al.
Diversification of photoperiodic response patterns in a collection of early-flowering mutants of Arabidopsis.
Plant Physiol., 2008. 148(3): p. 1465-73
[PMID:18799658] - Reňák D,Dupl'áková N,Honys D
Wide-scale screening of T-DNA lines for transcription factor genes affecting male gametophyte development in Arabidopsis.
Sex. Plant Reprod., 2012. 25(1): p. 39-60
[PMID:22101548] - Gu X,Wang Y,He Y
Photoperiodic regulation of flowering time through periodic histone deacetylation of the florigen gene FT.
PLoS Biol., 2013. 11(9): p. e1001649
[PMID:24019760] - Fernandez DE, et al.
The MADS-Domain Factors AGAMOUS-LIKE15 and AGAMOUS-LIKE18, along with SHORT VEGETATIVE PHASE and AGAMOUS-LIKE24, Are Necessary to Block Floral Gene Expression during the Vegetative Phase.
Plant Physiol., 2014. 165(4): p. 1591-1603
[PMID:24948837] - Serivichyaswat P, et al.
Expression of the floral repressor miRNA156 is positively regulated by the AGAMOUS-like proteins AGL15 and AGL18.
Mol. Cells, 2015. 38(3): p. 259-66
[PMID:25666346] - Jin J, et al.
An Arabidopsis Transcriptional Regulatory Map Reveals Distinct Functional and Evolutionary Features of Novel Transcription Factors.
Mol. Biol. Evol., 2015. 32(7): p. 1767-73
[PMID:25750178]
|




