| Signature Domain? help Back to Top |
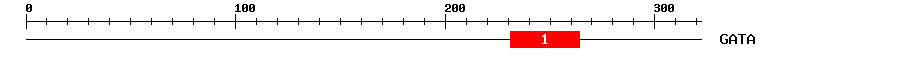 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | GATA | 56.5 | 3.9e-18 | 231 | 264 | 1 | 34 |
GATA 1 CsnCgttkTplWRrgpdgnktLCnaCGlyyrkkg 34
C +C +tkTp+WR gp g+ktLCnaCG++y++ +
AT3G54810.2 231 CMHCEVTKTPQWRLGPMGPKTLCNACGVRYKSGR 264
99*****************************987 PP
|
| Publications
? help Back to Top |
- Riechmann JL, et al.
Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes.
Science, 2000. 290(5499): p. 2105-10
[PMID:11118137] - Gutierrez RA,Ewing RM,Cherry JM,Green PJ
Identification of unstable transcripts in Arabidopsis by cDNA microarray analysis: rapid decay is associated with a group of touch- and specific clock-controlled genes.
Proc. Natl. Acad. Sci. U.S.A., 2002. 99(17): p. 11513-8
[PMID:12167669] - Jeong MJ,Jeong MJ,Shih MC
Interaction of a GATA factor with cis-acting elements involved in light regulation of nuclear genes encoding chloroplast glyceraldehyde-3-phosphate dehydrogenase in Arabidopsis.
Biochem. Biophys. Res. Commun., 2003. 300(2): p. 555-62
[PMID:12504119] - Mittelsten Scheid O,Afsar K,Paszkowski J
Formation of stable epialleles and their paramutation-like interaction in tetraploid Arabidopsis thaliana.
Nat. Genet., 2003. 34(4): p. 450-4
[PMID:12847525] - Yamada K, et al.
Empirical analysis of transcriptional activity in the Arabidopsis genome.
Science, 2003. 302(5646): p. 842-6
[PMID:14593172] - Reyes JC,Muro-Pastor MI,Florencio FJ
The GATA family of transcription factors in Arabidopsis and rice.
Plant Physiol., 2004. 134(4): p. 1718-32
[PMID:15084732] - Guan Y,Nothnagel EA
Binding of arabinogalactan proteins by Yariv phenylglycoside triggers wound-like responses in Arabidopsis cell cultures.
Plant Physiol., 2004. 135(3): p. 1346-66
[PMID:15235117] - Lu Y,Zhu J,Liu P
A two-step strategy for detecting differential gene expression in cDNA microarray data.
Curr. Genet., 2005. 47(2): p. 121-31
[PMID:15688252] - Bi YM, et al.
Genetic analysis of Arabidopsis GATA transcription factor gene family reveals a nitrate-inducible member important for chlorophyll synthesis and glucose sensitivity.
Plant J., 2005. 44(4): p. 680-92
[PMID:16262716] - Liu PP,Koizuka N,Martin RC,Nonogaki H
The BME3 (Blue Micropylar End 3) GATA zinc finger transcription factor is a positive regulator of Arabidopsis seed germination.
Plant J., 2005. 44(6): p. 960-71
[PMID:16359389] - Manfield IW,Devlin PF,Jen CH,Westhead DR,Gilmartin PM
Conservation, convergence, and divergence of light-responsive, circadian-regulated, and tissue-specific expression patterns during evolution of the Arabidopsis GATA gene family.
Plant Physiol., 2007. 143(2): p. 941-58
[PMID:17208962] - Lee J, et al.
Analysis of transcription factor HY5 genomic binding sites revealed its hierarchical role in light regulation of development.
Plant Cell, 2007. 19(3): p. 731-49
[PMID:17337630] - Sottosanto JB,Saranga Y,Blumwald E
Impact of AtNHX1, a vacuolar Na+/H+ antiporter, upon gene expression during short- and long-term salt stress in Arabidopsis thaliana.
BMC Plant Biol., 2007. 7: p. 18
[PMID:17411438] - Goeres DC, et al.
Components of the Arabidopsis mRNA decapping complex are required for early seedling development.
Plant Cell, 2007. 19(5): p. 1549-64
[PMID:17513503] - Choi D, et al.
iNID: an analytical framework for identifying network models for interplays among developmental signaling in Arabidopsis.
Mol Plant, 2014. 7(5): p. 792-813
[PMID:24380880] - Li T,Wu XY,Li H,Song JH,Liu JY
A Dual-Function Transcription Factor, AtYY1, Is a Novel Negative Regulator of the Arabidopsis ABA Response Network.
Mol Plant, 2016. 9(5): p. 650-661
[PMID:26961720]
|




