| Signature Domain? help Back to Top |
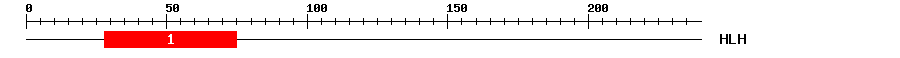 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | HLH | 14.7 | 5.5e-05 | 28 | 75 | 1 | 52 |
CHHHHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHH CS
HLH 1 rrrahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIk 52
+r ++ rEr R+++N+ f eL + l + ++ Ka+iL +A++++k
AT3G47640.2 28 KRINKAVRERLKREHLNELFIELADTLELN----QQNSGKASILCEATRFLK 75
689999*********************997....566778889999999887 PP
|
| Publications
? help Back to Top |
- Riechmann JL, et al.
Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes.
Science, 2000. 290(5499): p. 2105-10
[PMID:11118137] - Heim MA, et al.
The basic helix-loop-helix transcription factor family in plants: a genome-wide study of protein structure and functional diversity.
Mol. Biol. Evol., 2003. 20(5): p. 735-47
[PMID:12679534] - Toledo-Ortiz G,Huq E,Quail PH
The Arabidopsis basic/helix-loop-helix transcription factor family.
Plant Cell, 2003. 15(8): p. 1749-70
[PMID:12897250] - Yamada K, et al.
Empirical analysis of transcriptional activity in the Arabidopsis genome.
Science, 2003. 302(5646): p. 842-6
[PMID:14593172] - Bailey PC, et al.
Update on the basic helix-loop-helix transcription factor gene family in Arabidopsis thaliana.
Plant Cell, 2003. 15(11): p. 2497-502
[PMID:14600211] - Yang KY,Kim YM,Lee S,Song PS,Soh MS
Overexpression of a mutant basic helix-loop-helix protein HFR1, HFR1-deltaN105, activates a branch pathway of light signaling in Arabidopsis.
Plant Physiol., 2003. 133(4): p. 1630-42
[PMID:14645731] - Guan Y,Nothnagel EA
Binding of arabinogalactan proteins by Yariv phenylglycoside triggers wound-like responses in Arabidopsis cell cultures.
Plant Physiol., 2004. 135(3): p. 1346-66
[PMID:15235117] - Gong W, et al.
The development of protein microarrays and their applications in DNA-protein and protein-protein interaction analyses of Arabidopsis transcription factors.
Mol Plant, 2008. 1(1): p. 27-41
[PMID:19802365] - Long TA, et al.
The bHLH transcription factor POPEYE regulates response to iron deficiency in Arabidopsis roots.
Plant Cell, 2010. 22(7): p. 2219-36
[PMID:20675571] - Golisz A,Sugano M,Hiradate S,Fujii Y
Microarray analysis of Arabidopsis plants in response to allelochemical L-DOPA.
Planta, 2011. 233(2): p. 231-40
[PMID:20978802] - Arabidopsis Interactome Mapping Consortium
Evidence for network evolution in an Arabidopsis interactome map.
Science, 2011. 333(6042): p. 601-7
[PMID:21798944] - Ivanov R,Brumbarova T,Bauer P
Fitting into the harsh reality: regulation of iron-deficiency responses in dicotyledonous plants.
Mol Plant, 2012. 5(1): p. 27-42
[PMID:21873619] - Efroni I, et al.
Regulation of leaf maturation by chromatin-mediated modulation of cytokinin responses.
Dev. Cell, 2013. 24(4): p. 438-45
[PMID:23449474] - Rodríguez-Celma J, et al.
The transcriptional response of Arabidopsis leaves to Fe deficiency.
Front Plant Sci, 2013. 4: p. 276
[PMID:23888164] - Ding Y, et al.
Four distinct types of dehydration stress memory genes in Arabidopsis thaliana.
BMC Plant Biol., 2013. 13: p. 229
[PMID:24377444] - Martínez-Trujillo M, et al.
Chromate alters root system architecture and activates expression of genes involved in iron homeostasis and signaling in Arabidopsis thaliana.
Plant Mol. Biol., 2014. 86(1-2): p. 35-50
[PMID:24928490] - Li H,Wang L,Yang ZM
Co-expression analysis reveals a group of genes potentially involved in regulation of plant response to iron-deficiency.
Gene, 2015. 554(1): p. 16-24
[PMID:25300251] - Selote D,Samira R,Matthiadis A,Gillikin JW,Long TA
Iron-binding E3 ligase mediates iron response in plants by targeting basic helix-loop-helix transcription factors.
Plant Physiol., 2015. 167(1): p. 273-86
[PMID:25452667] - Liang G,Zhang H,Li X,Ai Q,Yu D
bHLH transcription factor bHLH115 regulates iron homeostasis in Arabidopsis thaliana.
J. Exp. Bot., 2017. 68(7): p. 1743-1755
[PMID:28369511] - Min JH, et al.
Arabidopsis Basic Helix-Loop-Helix 34 (bHLH34) Is Involved in Glucose Signaling through Binding to a GAGA Cis-Element.
Front Plant Sci, 2017. 8: p. 2100
[PMID:29321786]
|




