 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT3G46070.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 170aa MW: 18754.3 Da PI: 8.8553 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 14.9 | 7.9e-05 | 86 | 108 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
++Cp+Cg F+ L H+r+H
AT3G46070.1 86 HTCPICGLEFPMGQALGGHMRKH 108
79********************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF57667 | 2.16E-7 | 34 | 108 | No hit | No description |
| Pfam | PF13912 | 1.2E-9 | 35 | 60 | IPR007087 | Zinc finger, C2H2 |
| PROSITE profile | PS50157 | 9.12 | 36 | 63 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 2.1 | 36 | 58 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 38 | 58 | IPR007087 | Zinc finger, C2H2 |
| Pfam | PF13912 | 9.8E-11 | 85 | 109 | IPR007087 | Zinc finger, C2H2 |
| PROSITE profile | PS50157 | 10.824 | 86 | 113 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.034 | 86 | 108 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 88 | 108 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009046 | anatomy | flower | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 170 aa Download sequence Send to blast |
MVAESDNRDL TVDTAASCLM LLSGIGEHDG RKKRVFRCKT CERDFDSFQA LGGHRASHSK 60 LTNSDDKSLP GSPKKKPKTT TTTTAHTCPI CGLEFPMGQA LGGHMRKHRN EKEREKASNV 120 LVTHSFMPET TTVTTLKKSS SGKRVACLDF DLTSVESFVN TELELGRTMY |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 72 | 78 | PKKKPKT |
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| Genevisible | 252567_at | 0.0 | ||||
| Expression Atlas | AT3G46070 | - | ||||
| AtGenExpress | AT3G46070 | - | ||||
| ATTED-II | AT3G46070 | - | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may be involved in stress responses. {ECO:0000250}. | |||||
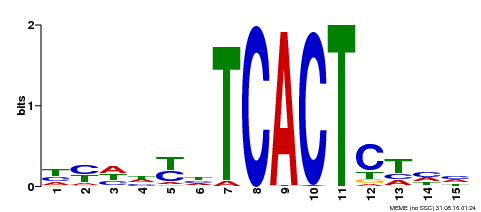
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00393 | DAP | 27203113 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT3G46070.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By heat stress. {ECO:0000269|PubMed:18201973}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT3G46070 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AL355775 | 0.0 | AL355775.1 Arabidopsis thaliana DNA chromosome 3, BAC clone F12M12. | |||
| GenBank | CP002686 | 0.0 | CP002686.1 Arabidopsis thaliana chromosome 3, complete sequence. | |||
| GenBank | DQ446739 | 0.0 | DQ446739.1 Arabidopsis thaliana clone pENTR221-At3g46070 zinc finger family protein (At3g46070) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_190193.1 | 1e-123 | C2H2-type zinc finger family protein | ||||
| Swissprot | Q9LX85 | 6e-71 | ZAT8_ARATH; Zinc finger protein ZAT8 | ||||
| TrEMBL | A0MF10 | 1e-122 | A0MF10_ARATH; Uncharacterized protein (Fragment) | ||||
| TrEMBL | Q9LX86 | 1e-122 | Q9LX86_ARATH; C2H2-type zinc finger family protein | ||||
| STRING | AT3G46070.1 | 1e-123 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2028 | 18 | 76 | Representative plant | OGRP131 | 15 | 149 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT3G46070.1 |
| Entrez Gene | 823750 |
| iHOP | AT3G46070 |
| wikigenes | AT3G46070 |



