 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT3G25730.1 | ||||||||
| Common Name | EDF3, K13N2.14, K13N2.7 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | RAV | ||||||||
| Protein Properties | Length: 333aa MW: 37755.8 Da PI: 9.7218 | ||||||||
| Description | ethylene response DNA binding factor 3 | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 40.6 | 6.5e-13 | 60 | 108 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s++kGV + +grW A+I++ ++r++lg+f ++eAa+a++ a+ +++g
AT3G25730.1 60 SRFKGVVPQP-NGRWGAQIYEK-----HQRVWLGTFNEEDEAARAYDVAAHRFRG 108
689***9788.8*********3.....5**********99************998 PP
| |||||||
| 2 | B3 | 102 | 3.3e-32 | 183 | 288 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE. CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefe 91
f+k+ tpsdv+k++rlv+pk+ ae+h ++++ +++ l +ed +g++W+++++y+++s++yvltkGW++Fvk+++L++gD + Fk++++++++
AT3G25730.1 183 FEKTVTPSDVGKLNRLVIPKHQAEKHfplplgnNNVSVKGMLLNFEDVNGKVWRFRYSYWNSSQSYVLTKGWSRFVKEKRLCAGDLISFKRSNDQDQK 280
99*****************************9977888****************************************************99999999 PP
.EEEEE-S CS
B3 92 lvvkvfrk 99
+++++ +k
AT3G25730.1 281 FFIGWKSK 288
99999876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF00847 | 2.9E-8 | 60 | 108 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 5.79E-26 | 60 | 118 | No hit | No description |
| SuperFamily | SSF54171 | 3.66E-17 | 60 | 117 | IPR016177 | DNA-binding domain |
| Gene3D | G3DSA:3.30.730.10 | 2.2E-20 | 61 | 117 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 3.8E-28 | 61 | 122 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 20.086 | 61 | 116 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:2.40.330.10 | 5.2E-39 | 176 | 289 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 1.96E-31 | 181 | 290 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 7.54E-27 | 182 | 276 | No hit | No description |
| Pfam | PF02362 | 2.9E-28 | 183 | 288 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 13.987 | 183 | 289 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 3.6E-25 | 183 | 289 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000293 | anatomy | guard cell | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009010 | anatomy | seed | ||||
| PO:0009046 | anatomy | flower | ||||
| PO:0020030 | anatomy | cotyledon | ||||
| PO:0020100 | anatomy | hypocotyl | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 333 aa Download sequence Send to blast |
MDAMSSVDES STTTDSIPAR KSSSPASLLY RMGSGTSVVL DSENGVEVEV EAESRKLPSS 60 RFKGVVPQPN GRWGAQIYEK HQRVWLGTFN EEDEAARAYD VAAHRFRGRD AVTNFKDTTF 120 EEEVEFLNAH SKSEIVDMLR KHTYKEELDQ RKRNRDGNGK ETTAFALASM VVMTGFKTAE 180 LLFEKTVTPS DVGKLNRLVI PKHQAEKHFP LPLGNNNVSV KGMLLNFEDV NGKVWRFRYS 240 YWNSSQSYVL TKGWSRFVKE KRLCAGDLIS FKRSNDQDQK FFIGWKSKSG LDLETGRVMR 300 LFGVDISLNA VVVVKETTEV LMSSLRCKKQ RVL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wid_A | 9e-65 | 179 | 295 | 10 | 124 | DNA-binding protein RAV1 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| At.37331 | 0.0 | seed | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 186510423 | 0.0 | ||||
| Genevisible | 257643_at | 0.0 | ||||
| Expression Atlas | AT3G25730 | - | ||||
| AtGenExpress | AT3G25730 | - | ||||
| ATTED-II | AT3G25730 | - | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probably acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. May be involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways (By similarity). {ECO:0000250}. | |||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
|
||||||
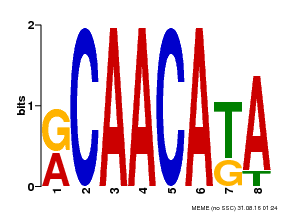
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00031 | PBM | 25215497 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT3G25730.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT3G25730 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB028607 | 0.0 | AB028607.1 Arabidopsis thaliana genomic DNA, chromosome 3, TAC clone:K13N2. | |||
| GenBank | AJ441073 | 0.0 | AJ441073.1 Arabidopsis thaliana mRNA for putative auxin response factor 14 (arf14 gene). | |||
| GenBank | AK229415 | 0.0 | AK229415.1 Arabidopsis thaliana mRNA for AP2 domain transcription factor, complete cds, clone: RAFL16-68-B16. | |||
| GenBank | CP002686 | 0.0 | CP002686.1 Arabidopsis thaliana chromosome 3, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_189201.1 | 0.0 | ethylene response DNA binding factor 3 | ||||
| Swissprot | Q9LS06 | 0.0 | RAVL4_ARATH; AP2/ERF and B3 domain-containing transcription factor ARF14 | ||||
| TrEMBL | A0A178VJD4 | 0.0 | A0A178VJD4_ARATH; EDF3 | ||||
| STRING | AT3G25730.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM848 | 27 | 121 | Representative plant | OGRP365 | 16 | 108 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT3G25730.1 |
| Entrez Gene | 822164 |
| iHOP | AT3G25730 |
| wikigenes | AT3G25730 |



