| Signature Domain? help Back to Top |
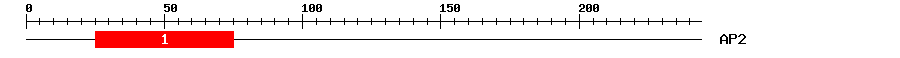 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | AP2 | 57.7 | 2.8e-18 | 25 | 75 | 1 | 55 |
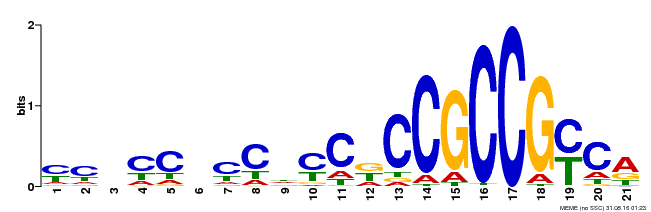
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
++y+GVr ++ +gr++AeIrdp + + r++lg+f++a +Aa+a++ a+++l+g
AT3G20310.1 25 PRYRGVRKRP-WGRFAAEIRDPLK---KSRVWLGTFDSAVDAARAYDTAARNLRG 75
68********.**********743...5*************************99 PP
|
| Annotation --
Nucleotide ? help
Back to Top |
| Source |
Hit ID |
E-value |
Description |
| GenBank | AB024036 | 0.0 | AB024036.1 Arabidopsis thaliana genomic DNA, chromosome 3, P1 clone: MQC12. |
| GenBank | AB032201 | 0.0 | AB032201.1 Arabidopsis thaliana mRNA for ERF transcription factor 7, complete cds. |
| GenBank | AK228460 | 0.0 | AK228460.1 Arabidopsis thaliana mRNA for putative ethylene responsive element binding factor, complete cds, clone: RAFL15-05-J12. |
| GenBank | AY037254 | 0.0 | AY037254.1 Arabidopsis thaliana AT3g20310/MQC12_6 mRNA, complete cds. |
| GenBank | AY094001 | 0.0 | AY094001.1 Arabidopsis thaliana AT3g20310/MQC12_6 mRNA, complete cds. |
| GenBank | CP002686 | 0.0 | CP002686.1 Arabidopsis thaliana chromosome 3, complete sequence. |
| Publications
? help Back to Top |
- Riechmann JL, et al.
Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes.
Science, 2000. 290(5499): p. 2105-10
[PMID:11118137] - Ohta M,Matsui K,Hiratsu K,Shinshi H,Ohme-Takagi M
Repression domains of class II ERF transcriptional repressors share an essential motif for active repression.
Plant Cell, 2001. 13(8): p. 1959-68
[PMID:11487705] - Yamada K, et al.
Empirical analysis of transcriptional activity in the Arabidopsis genome.
Science, 2003. 302(5646): p. 842-6
[PMID:14593172] - Cluis CP,Mouchel CF,Hardtke CS
The Arabidopsis transcription factor HY5 integrates light and hormone signaling pathways.
Plant J., 2004. 38(2): p. 332-47
[PMID:15078335] - Song CP, et al.
Role of an Arabidopsis AP2/EREBP-type transcriptional repressor in abscisic acid and drought stress responses.
Plant Cell, 2005. 17(8): p. 2384-96
[PMID:15994908] - Duarte JM, et al.
Expression pattern shifts following duplication indicative of subfunctionalization and neofunctionalization in regulatory genes of Arabidopsis.
Mol. Biol. Evol., 2006. 23(2): p. 469-78
[PMID:16280546] - Babula D, et al.
Genes involved in biosynthesis and signalisation of ethylene in Brassica oleracea and Arabidopsis thaliana: identification and genome comparative mapping of specific gene homologues.
Theor. Appl. Genet., 2006. 112(3): p. 410-20
[PMID:16311726] - Nakano T,Suzuki K,Fujimura T,Shinshi H
Genome-wide analysis of the ERF gene family in Arabidopsis and rice.
Plant Physiol., 2006. 140(2): p. 411-32
[PMID:16407444] - Nemhauser JL,Hong F,Chory J
Different plant hormones regulate similar processes through largely nonoverlapping transcriptional responses.
Cell, 2006. 126(3): p. 467-75
[PMID:16901781] - Jung J, et al.
The barley ERF-type transcription factor HvRAF confers enhanced pathogen resistance and salt tolerance in Arabidopsis.
Planta, 2007. 225(3): p. 575-88
[PMID:16937017] - Manavella PA, et al.
Cross-talk between ethylene and drought signalling pathways is mediated by the sunflower Hahb-4 transcription factor.
Plant J., 2006. 48(1): p. 125-37
[PMID:16972869] - Galuschka C,Schindler M,Bülow L,Hehl R
AthaMap web tools for the analysis and identification of co-regulated genes.
Nucleic Acids Res., 2007. 35(Database issue): p. D857-62
[PMID:17148485] - Wang Y, et al.
Transcriptome analyses show changes in gene expression to accompany pollen germination and tube growth in Arabidopsis.
Plant Physiol., 2008. 148(3): p. 1201-11
[PMID:18775970] - Kagale S,Links MG,Rozwadowski K
Genome-wide analysis of ethylene-responsive element binding factor-associated amphiphilic repression motif-containing transcriptional regulators in Arabidopsis.
Plant Physiol., 2010. 152(3): p. 1109-34
[PMID:20097792] - Arabidopsis Interactome Mapping Consortium
Evidence for network evolution in an Arabidopsis interactome map.
Science, 2011. 333(6042): p. 601-7
[PMID:21798944] - Klopffleisch K, et al.
Arabidopsis G-protein interactome reveals connections to cell wall carbohydrates and morphogenesis.
Mol. Syst. Biol., 2011. 7: p. 532
[PMID:21952135] - Gaudinier A, et al.
Enhanced Y1H assays for Arabidopsis.
Nat. Methods, 2011. 8(12): p. 1053-5
[PMID:22037706] - Causier B,Ashworth M,Guo W,Davies B
The TOPLESS interactome: a framework for gene repression in Arabidopsis.
Plant Physiol., 2012. 158(1): p. 423-38
[PMID:22065421] - Ding Y, et al.
Four distinct types of dehydration stress memory genes in Arabidopsis thaliana.
BMC Plant Biol., 2013. 13: p. 229
[PMID:24377444] - Jin J, et al.
An Arabidopsis Transcriptional Regulatory Map Reveals Distinct Functional and Evolutionary Features of Novel Transcription Factors.
Mol. Biol. Evol., 2015. 32(7): p. 1767-73
[PMID:25750178] - Giuntoli B,Perata P
Group VII Ethylene Response Factors in Arabidopsis: Regulation and Physiological Roles.
Plant Physiol., 2018. 176(2): p. 1143-1155
[PMID:29269576] - Riechmann JL,Meyerowitz EM
The AP2/EREBP family of plant transcription factors.
Biol. Chem., 1998. 379(6): p. 633-46
[PMID:9687012]
|