 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT3G03200.1 | ||||||||
| Common Name | anac045, NAC045, T17B22.11 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 479aa MW: 55587.6 Da PI: 5.3407 | ||||||||
| Description | NAC domain containing protein 45 | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 176 | 1.1e-54 | 6 | 133 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLp.k.kvkaeekewyfFskrdkkyatgkrknratksgyWkatgkdkevlskk 96
lppGfrFhPtdeel+++yLk+k++g ++el evi+evd+yk+ePwdLp k + ++++ewyfFs+rd+ky++g+r+nratk gyWkatgkd++v +
AT3G03200.1 6 LPPGFRFHPTDEELITYYLKRKINGLEIEL-EVIAEVDLYKCEPWDLPgKsLLPSKDQEWYFFSPRDRKYPNGSRTNRATKGGYWKATGKDRRVSW-R 101
79****************************.99***************6345667999*************************************9.8 PP
NAM 97 gelvglkktLvfykgrapkgektdWvmheyrl 128
++++g+kktLv+y+grap+g +t Wvmheyrl
AT3G03200.1 102 DRAIGTKKTLVYYRGRAPHGIRTGWVMHEYRL 133
899***************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 9.81E-63 | 4 | 157 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 61.025 | 6 | 157 | IPR003441 | NAC domain |
| Pfam | PF02365 | 7.1E-29 | 7 | 133 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007275 | Biological Process | multicellular organism development | ||||
| GO:0090602 | Biological Process | sieve element enucleation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009029 | anatomy | stamen | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 479 aa Download sequence Send to blast |
MAPVSLPPGF RFHPTDEELI TYYLKRKING LEIELEVIAE VDLYKCEPWD LPGKSLLPSK 60 DQEWYFFSPR DRKYPNGSRT NRATKGGYWK ATGKDRRVSW RDRAIGTKKT LVYYRGRAPH 120 GIRTGWVMHE YRLDETECEP SAYGMQDAYA LCRVFKKIVI EAKPRDQHRS YVHAMSNVSG 180 NCSSSFDTCS DLEISSTTHQ VQNTFQPRFG NERFNSNAIS NEDWSQYYGS SYRPFPTPYK 240 VNTEIECSML QHNIYLPPLR VENSAFSDSD FFTSMTHNND HGVFDDFTFA ASNSNHNNSV 300 GDQVIHVGNY DEQLITSNRH MNQTGYIKEQ KIRSSLDNTD EDPGFHGNNT NDNIDIDDFL 360 SFDIYNEDNV NQIEDNEDVN TNETLDSSGF EVVEEETRFN NQMLISTYQT TKILYHQVVP 420 CHTLKVHVNP ISHNVEERTL FIEEDKDSWL QRAEKITKTK LTLFSLMAQQ YYKCLAIFF |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ut4_A | 3e-55 | 1 | 157 | 12 | 165 | NO APICAL MERISTEM PROTEIN |
| 1ut4_B | 3e-55 | 1 | 157 | 12 | 165 | NO APICAL MERISTEM PROTEIN |
| 1ut7_A | 3e-55 | 1 | 157 | 12 | 165 | NO APICAL MERISTEM PROTEIN |
| 1ut7_B | 3e-55 | 1 | 157 | 12 | 165 | NO APICAL MERISTEM PROTEIN |
| 4dul_A | 3e-55 | 1 | 157 | 12 | 165 | NAC domain-containing protein 19 |
| 4dul_B | 3e-55 | 1 | 157 | 12 | 165 | NAC domain-containing protein 19 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| At.41017 | 0.0 | leaf | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 186509709 | 0.0 | ||||
| Genevisible | 258865_at | 0.0 | ||||
| Expression Atlas | AT3G03200 | - | ||||
| AtGenExpress | AT3G03200 | - | ||||
| ATTED-II | AT3G03200 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in a few sieve element cells before enucleation and in phloem-pole pericycle cells. {ECO:0000269|PubMed:25081480}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor directing sieve element enucleation and cytosol degradation. Not required for formation of lytic vacuoles. Regulates, with NAC086, the transcription of NEN1, NEN2, NEN3, NEN4, RTM1, RTM2, UBP16, PLDZETA, ABCB10 and At1g26450. {ECO:0000269|PubMed:25081480}. | |||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
|
||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00329 | DAP | 27203113 | Download |
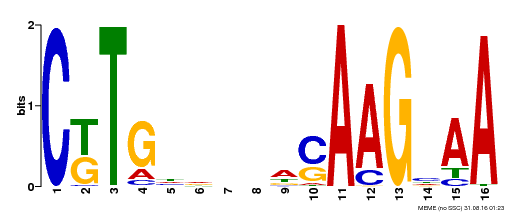
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT3G03200.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Regulated by the transcription factor APL (AC Q9SAK5). {ECO:0000269|PubMed:25081480}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Interaction ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Intact With | |||||
| IntAct | Search A4VCM0 | |||||
| Phenotype -- Disruption Phenotype ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | DISRUPTION PHENOTYPE: No visible phenotype, due to the redundancy with NAC086 (AC Q9FFI5). Nac045 and nac086 double mutants exhibit seedling lethality with defective development of the protophloem sieve element. {ECO:0000269|PubMed:25081480}. | |||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT3G03200 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT030461 | 0.0 | BT030461.1 Arabidopsis thaliana At3g03200 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_186970.2 | 0.0 | NAC domain containing protein 45 | ||||
| Swissprot | A4VCM0 | 0.0 | NAC45_ARATH; NAC domain-containing protein 45 | ||||
| TrEMBL | A0A178VIZ2 | 0.0 | A0A178VIZ2_ARATH; NAC045 | ||||
| STRING | AT3G03200.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4411 | 18 | 52 | Representative plant | OGRP17 | 15 | 800 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT3G03200.1 |
| Entrez Gene | 821226 |
| iHOP | AT3G03200 |
| wikigenes | AT3G03200 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



