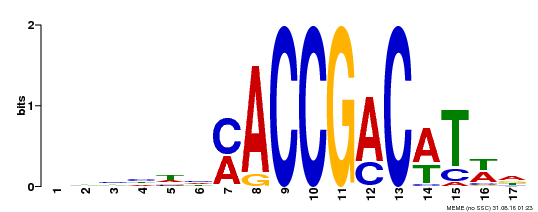
| Signature Domain? help Back to Top |
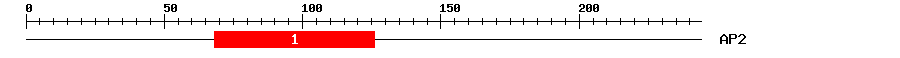 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | AP2 | 59.1 | 1e-18 | 68 | 126 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpse.ng....krkrfslgkfgtaeeAakaaiaarkkleg 55
++++GVr++ +g+WvAeIr+p++ +g ++kr +lg+f ta eAa a+++a+ ++g
AT2G38340.1 68 CRFRGVRQRV-WGKWVAEIREPVShRGanssRSKRLWLGTFATAAEAALAYDRAASVMYG 126
689*****99.**********888777877555*********************988776 PP
|
| Publications
? help Back to Top |
- Riechmann JL, et al.
Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes.
Science, 2000. 290(5499): p. 2105-10
[PMID:11118137] - Sakuma Y, et al.
DNA-binding specificity of the ERF/AP2 domain of Arabidopsis DREBs, transcription factors involved in dehydration- and cold-inducible gene expression.
Biochem. Biophys. Res. Commun., 2002. 290(3): p. 998-1009
[PMID:11798174] - Dal Bosco C, et al.
Inactivation of the chloroplast ATP synthase gamma subunit results in high non-photochemical fluorescence quenching and altered nuclear gene expression in Arabidopsis thaliana.
J. Biol. Chem., 2004. 279(2): p. 1060-9
[PMID:14576160] - Yamada K, et al.
Empirical analysis of transcriptional activity in the Arabidopsis genome.
Science, 2003. 302(5646): p. 842-6
[PMID:14593172] - Scheible WR, et al.
Genome-wide reprogramming of primary and secondary metabolism, protein synthesis, cellular growth processes, and the regulatory infrastructure of Arabidopsis in response to nitrogen.
Plant Physiol., 2004. 136(1): p. 2483-99
[PMID:15375205] - Nakano T,Suzuki K,Fujimura T,Shinshi H
Genome-wide analysis of the ERF gene family in Arabidopsis and rice.
Plant Physiol., 2006. 140(2): p. 411-32
[PMID:16407444] - Truman W,de Zabala MT,Grant M
Type III effectors orchestrate a complex interplay between transcriptional networks to modify basal defence responses during pathogenesis and resistance.
Plant J., 2006. 46(1): p. 14-33
[PMID:16553893] - Che P,Lall S,Nettleton D,Howell SH
Gene expression programs during shoot, root, and callus development in Arabidopsis tissue culture.
Plant Physiol., 2006. 141(2): p. 620-37
[PMID:16648215] - Mandaokar A, et al.
Transcriptional regulators of stamen development in Arabidopsis identified by transcriptional profiling.
Plant J., 2006. 46(6): p. 984-1008
[PMID:16805732] - Michel K,Abderhalden O,Bruggmann R,Dudler R
Transcriptional changes in powdery mildew infected wheat and Arabidopsis leaves undergoing syringolin-triggered hypersensitive cell death at infection sites.
Plant Mol. Biol., 2006. 62(4-5): p. 561-78
[PMID:16941219] - Sattler SE, et al.
Nonenzymatic lipid peroxidation reprograms gene expression and activates defense markers in Arabidopsis tocopherol-deficient mutants.
Plant Cell, 2006. 18(12): p. 3706-20
[PMID:17194769] - Culligan KM,Robertson CE,Foreman J,Doerner P,Britt AB
ATR and ATM play both distinct and additive roles in response to ionizing radiation.
Plant J., 2006. 48(6): p. 947-61
[PMID:17227549] - Veyres N, et al.
The Arabidopsis sweetie mutant is affected in carbohydrate metabolism and defective in the control of growth, development and senescence.
Plant J., 2008. 55(4): p. 665-86
[PMID:18452589] - Welling A,Palva ET
Involvement of CBF transcription factors in winter hardiness in birch.
Plant Physiol., 2008. 147(3): p. 1199-211
[PMID:18467468] - Ascencio-Ib
Global analysis of Arabidopsis gene expression uncovers a complex array of changes impacting pathogen response and cell cycle during geminivirus infection.
Plant Physiol., 2008. 148(1): p. 436-54
[PMID:18650403] - Wang Y, et al.
Transcriptome analyses show changes in gene expression to accompany pollen germination and tube growth in Arabidopsis.
Plant Physiol., 2008. 148(3): p. 1201-11
[PMID:18775970] - Krishnaswamy SS, et al.
Transcriptional profiling of pea ABR17 mediated changes in gene expression in Arabidopsis thaliana.
BMC Plant Biol., 2008. 8: p. 91
[PMID:18783601] - Dubos C, et al.
MYB transcription factors in Arabidopsis.
Trends Plant Sci., 2010. 15(10): p. 573-81
[PMID:20674465] - Krishnaswamy S,Verma S,Rahman MH,Kav NN
Functional characterization of four APETALA2-family genes (RAP2.6, RAP2.6L, DREB19 and DREB26) in Arabidopsis.
Plant Mol. Biol., 2011. 75(1-2): p. 107-27
[PMID:21069430] - Gaudinier A, et al.
Enhanced Y1H assays for Arabidopsis.
Nat. Methods, 2011. 8(12): p. 1053-5
[PMID:22037706] - Mehterov N, et al.
Oxidative stress provokes distinct transcriptional responses in the stress-tolerant atr7 and stress-sensitive loh2 Arabidopsis thaliana mutants as revealed by multi-parallel quantitative real-time PCR analysis of ROS marker and antioxidant genes.
Plant Physiol. Biochem., 2012. 59: p. 20-9
[PMID:22710144] - Huang KC,Lin WC,Cheng WH
Salt hypersensitive mutant 9, a nucleolar APUM23 protein, is essential for salt sensitivity in association with the ABA signaling pathway in Arabidopsis.
BMC Plant Biol., 2018. 18(1): p. 40
[PMID:29490615]
|





