| Signature Domain? help Back to Top |
 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | SRF-TF | 89.5 | 1.8e-28 | 10 | 59 | 2 | 51 |
---SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-TTSEEEEEE- CS
SRF-TF 2 rienksnrqvtfskRrngilKKAeELSvLCdaevaviifsstgklyeyss 51
ri+n++ rqvtfskRr g+lKKA+ELS+LCdaev viifsstgkly+y+s
AT2G14210.1 10 RIDNSTSRQVTFSKRRSGLLKKAKELSILCDAEVGVIIFSSTGKLYDYAS 59
8***********************************************86 PP
|
| 2 | K-box | 82.7 | 8.2e-28 | 82 | 170 | 9 | 97 |
K-box 9 leeakaeslqqelakLkkeienLqreqRhllGedLesLslkeLqqLeqqLekslkkiRskKnellleqieelqkkekelqeenkaLrkk 97
+++++ + +q+e+a L+++++ Lq+ +R+l+Ge+L+ ++ ++Lq+Le+qL +slk +R kK++l++++i+el +k + +q+en++L++
AT2G14210.1 82 NHASEIKFWQREVASLQQQLQYLQECHRKLVGEELSGMNANDLQNLEDQLVTSLKGVRLKKDQLMTNEIRELNRKGQIIQKENHELQNI 170
688999********************************************************************************975 PP
|
| Protein Features
? help Back to Top |
 |
| Database |
Entry ID |
E-value |
Start |
End |
InterPro ID |
Description |
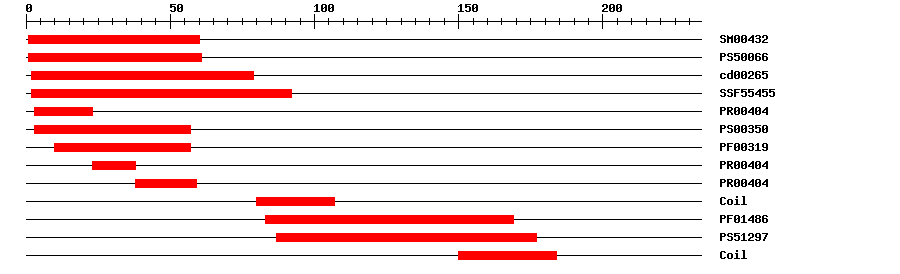
| SMART | SM00432 | 8.0E-40 | 1 | 60 | IPR002100 | Transcription factor, MADS-box |
| PROSITE profile | PS50066 | 31.665 | 1 | 61 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00265 | 1.90E-41 | 2 | 79 | No hit | No description |
| SuperFamily | SSF55455 | 2.22E-32 | 2 | 92 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 1.8E-28 | 3 | 23 | IPR002100 | Transcription factor, MADS-box |
| PROSITE pattern | PS00350 | 0 | 3 | 57 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF00319 | 1.9E-25 | 10 | 57 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 1.8E-28 | 23 | 38 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 1.8E-28 | 38 | 59 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF01486 | 5.8E-24 | 83 | 169 | IPR002487 | Transcription factor, K-box |
| PROSITE profile | PS51297 | 14.135 | 87 | 177 | IPR002487 | Transcription factor, K-box |
| Publications
? help Back to Top |
- Zhang H,Forde BG
Regulation of Arabidopsis root development by nitrate availability.
J. Exp. Bot., 2000. 51(342): p. 51-9
[PMID:10938795] - Wang R,Guegler K,LaBrie ST,Crawford NM
Genomic analysis of a nutrient response in Arabidopsis reveals diverse expression patterns and novel metabolic and potential regulatory genes induced by nitrate.
Plant Cell, 2000. 12(8): p. 1491-509
[PMID:10948265] - Riechmann JL, et al.
Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes.
Science, 2000. 290(5499): p. 2105-10
[PMID:11118137] - Parenicová L, et al.
Molecular and phylogenetic analyses of the complete MADS-box transcription factor family in Arabidopsis: new openings to the MADS world.
Plant Cell, 2003. 15(7): p. 1538-51
[PMID:12837945] - Yamada K, et al.
Empirical analysis of transcriptional activity in the Arabidopsis genome.
Science, 2003. 302(5646): p. 842-6
[PMID:14593172] - Filleur S,Walch-Liu P,Gan Y,Forde BG
Nitrate and glutamate sensing by plant roots.
Biochem. Soc. Trans., 2005. 33(Pt 1): p. 283-6
[PMID:15667327] - de Folter S, et al.
Comprehensive interaction map of the Arabidopsis MADS Box transcription factors.
Plant Cell, 2005. 17(5): p. 1424-33
[PMID:15805477] - Gan Y,Filleur S,Rahman A,Gotensparre S,Forde BG
Nutritional regulation of ANR1 and other root-expressed MADS-box genes in Arabidopsis thaliana.
Planta, 2005. 222(4): p. 730-42
[PMID:16021502] - Walch-Liu P, et al.
Nitrogen regulation of root branching.
Ann. Bot., 2006. 97(5): p. 875-81
[PMID:16339770] - De Bodt S,Theissen G,Van de Peer Y
Promoter analysis of MADS-box genes in eudicots through phylogenetic footprinting.
Mol. Biol. Evol., 2006. 23(6): p. 1293-303
[PMID:16581940] - Leseberg CH,Li A,Kang H,Duvall M,Mao L
Genome-wide analysis of the MADS-box gene family in Populus trichocarpa.
Gene, 2006. 378: p. 84-94
[PMID:16831523] - Remans T, et al.
The Arabidopsis NRT1.1 transporter participates in the signaling pathway triggering root colonization of nitrate-rich patches.
Proc. Natl. Acad. Sci. U.S.A., 2006. 103(50): p. 19206-11
[PMID:17148611] - Zhang H,Rong H,Pilbeam D
Signalling mechanisms underlying the morphological responses of the root system to nitrogen in Arabidopsis thaliana.
J. Exp. Bot., 2007. 58(9): p. 2329-38
[PMID:17578866] - Walch-Liu P,Forde BG
Nitrate signalling mediated by the NRT1.1 nitrate transporter antagonises L-glutamate-induced changes in root architecture.
Plant J., 2008. 54(5): p. 820-8
[PMID:18266918] - Castaings L, et al.
The nodule inception-like protein 7 modulates nitrate sensing and metabolism in Arabidopsis.
Plant J., 2009. 57(3): p. 426-35
[PMID:18826430] - Feng H, et al.
Multiple roles of nitrate transport accessory protein NAR2 in plants.
Plant Signal Behav, 2011. 6(9): p. 1286-9
[PMID:21852757] - Gan Y,Bernreiter A,Filleur S,Abram B,Forde BG
Overexpressing the ANR1 MADS-box gene in transgenic plants provides new insights into its role in the nitrate regulation of root development.
Plant Cell Physiol., 2012. 53(6): p. 1003-16
[PMID:22523192] - Heyndrickx KS,Vandepoele K
Systematic identification of functional plant modules through the integration of complementary data sources.
Plant Physiol., 2012. 159(3): p. 884-901
[PMID:22589469] - Yan Y,Wang H,Hamera S,Chen X,Fang R
miR444a has multiple functions in the rice nitrate-signaling pathway.
Plant J., 2014. 78(1): p. 44-55
[PMID:24460537] - Jin J, et al.
An Arabidopsis Transcriptional Regulatory Map Reveals Distinct Functional and Evolutionary Features of Novel Transcription Factors.
Mol. Biol. Evol., 2015. 32(7): p. 1767-73
[PMID:25750178] - Lei L, et al.
Nitrogen use efficiency is regulated by interacting proteins relevant to development in wheat.
Plant Biotechnol. J., 2018. 16(6): p. 1214-1226
[PMID:29193541] - Sun CH, et al.
Chrysanthemum MADS-box transcription factor CmANR1 modulates lateral root development via homo-/heterodimerization to influence auxin accumulation in Arabidopsis.
Plant Sci., 2018. 266: p. 27-36
[PMID:29241564] - Zhang H,Forde BG
An Arabidopsis MADS box gene that controls nutrient-induced changes in root architecture.
Science, 1998. 279(5349): p. 407-9
[PMID:9430595]
|




