 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT1G79580.2 | ||||||||
| Common Name | ANAC033, F20B17.1, NAC033, SMB, URP7 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 371aa MW: 42381.8 Da PI: 7.2243 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 169.9 | 8.2e-53 | 17 | 146 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLpk..kvka.eekewyfFskrdkkyatgkrknratksgyWkatgkdkevlsk 95
+ppGfrFhPt+eel+ +yLkkkv+ ++++l +vi+evd++k+ePw+L++ ++ + ++ewyfFs++dkky+tg+r+nrat++g+Wkatg+dk+++ +
AT1G79580.2 17 VPPGFRFHPTEEELLYYYLKKKVSYEPIDL-DVIREVDLNKLEPWELKEkcRIGSgPQNEWYFFSHKDKKYPTGTRTNRATAAGFWKATGRDKSIHLN 113
69****************************.9***************952444443456*************************************** PP
NAM 96 kgelvglkktLvfykgrapkgektdWvmheyrl 128
+++++gl+ktLvfy+grap+g+kt+W+mheyrl
AT1G79580.2 114 SSKKIGLRKTLVFYTGRAPHGQKTEWIMHEYRL 146
*******************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 9.94E-59 | 10 | 166 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 57.73 | 17 | 166 | IPR003441 | NAC domain |
| Pfam | PF02365 | 3.1E-27 | 18 | 146 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003002 | Biological Process | regionalization | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009834 | Biological Process | plant-type secondary cell wall biogenesis | ||||
| GO:0010455 | Biological Process | positive regulation of cell fate commitment | ||||
| GO:0048829 | Biological Process | root cap development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000013 | anatomy | cauline leaf | ||||
| PO:0000037 | anatomy | shoot apex | ||||
| PO:0000230 | anatomy | inflorescence meristem | ||||
| PO:0008019 | anatomy | leaf lamina base | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009009 | anatomy | plant embryo | ||||
| PO:0009010 | anatomy | seed | ||||
| PO:0009025 | anatomy | vascular leaf | ||||
| PO:0009029 | anatomy | stamen | ||||
| PO:0009030 | anatomy | carpel | ||||
| PO:0009031 | anatomy | sepal | ||||
| PO:0009032 | anatomy | petal | ||||
| PO:0009046 | anatomy | flower | ||||
| PO:0009047 | anatomy | stem | ||||
| PO:0009052 | anatomy | flower pedicel | ||||
| PO:0020030 | anatomy | cotyledon | ||||
| PO:0020038 | anatomy | petiole | ||||
| PO:0020100 | anatomy | hypocotyl | ||||
| PO:0020131 | anatomy | lateral root cap | ||||
| PO:0025022 | anatomy | collective leaf structure | ||||
| PO:0025281 | anatomy | pollen | ||||
| PO:0001054 | developmental stage | vascular leaf senescent stage | ||||
| PO:0001078 | developmental stage | plant embryo cotyledonary stage | ||||
| PO:0001081 | developmental stage | mature plant embryo stage | ||||
| PO:0001185 | developmental stage | plant embryo globular stage | ||||
| PO:0007064 | developmental stage | LP.12 twelve leaves visible stage | ||||
| PO:0007095 | developmental stage | LP.08 eight leaves visible stage | ||||
| PO:0007098 | developmental stage | LP.02 two leaves visible stage | ||||
| PO:0007103 | developmental stage | LP.10 ten leaves visible stage | ||||
| PO:0007115 | developmental stage | LP.04 four leaves visible stage | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| PO:0007616 | developmental stage | flowering stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 371 aa Download sequence Send to blast |
MEIGSSSTVA GGGQLSVPPG FRFHPTEEEL LYYYLKKKVS YEPIDLDVIR EVDLNKLEPW 60 ELKEKCRIGS GPQNEWYFFS HKDKKYPTGT RTNRATAAGF WKATGRDKSI HLNSSKKIGL 120 RKTLVFYTGR APHGQKTEWI MHEYRLDDSE NEIQEDGWVV CRVFKKKNHF RGFHQEQEQD 180 HHHHHQYIST NNDHDHHHHI DSNSNNHSPL ILHPLDHHHH HHHIGRQIHM PLHEFANTLS 240 HGSMHLPQLF SPDSAAAAAA AAASAQPFVS PINTTDIECS QNLLRLTSNN NYGGDWSFLD 300 KLLTTGNMNQ QQQQQVQNHQ AKCFGDLSNN DNNDQADHLG NNNGGSSSSP VNQRFPFHYL 360 GNDANLLKFP K |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ut4_A | 4e-50 | 14 | 168 | 14 | 167 | NO APICAL MERISTEM PROTEIN |
| 1ut4_B | 4e-50 | 14 | 168 | 14 | 167 | NO APICAL MERISTEM PROTEIN |
| 1ut7_A | 4e-50 | 14 | 168 | 14 | 167 | NO APICAL MERISTEM PROTEIN |
| 1ut7_B | 4e-50 | 14 | 168 | 14 | 167 | NO APICAL MERISTEM PROTEIN |
| 3swm_A | 5e-50 | 14 | 168 | 17 | 170 | NAC domain-containing protein 19 |
| 3swm_B | 5e-50 | 14 | 168 | 17 | 170 | NAC domain-containing protein 19 |
| 3swm_C | 5e-50 | 14 | 168 | 17 | 170 | NAC domain-containing protein 19 |
| 3swm_D | 5e-50 | 14 | 168 | 17 | 170 | NAC domain-containing protein 19 |
| 3swp_A | 5e-50 | 14 | 168 | 17 | 170 | NAC domain-containing protein 19 |
| 3swp_B | 5e-50 | 14 | 168 | 17 | 170 | NAC domain-containing protein 19 |
| 3swp_C | 5e-50 | 14 | 168 | 17 | 170 | NAC domain-containing protein 19 |
| 3swp_D | 5e-50 | 14 | 168 | 17 | 170 | NAC domain-containing protein 19 |
| 4dul_A | 4e-50 | 14 | 168 | 14 | 167 | NAC domain-containing protein 19 |
| 4dul_B | 4e-50 | 14 | 168 | 14 | 167 | NAC domain-containing protein 19 |
| Search in ModeBase | ||||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 42563353 | 0.0 | ||||
| Genevisible | 261393_at | 0.0 | ||||
| Expression Atlas | AT1G79580 | - | ||||
| AtGenExpress | AT1G79580 | - | ||||
| ATTED-II | AT1G79580 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: First expressed at early heart stage onward in all root basal daughter cells resulting from horizontal divisions in the COL progenitors and is later maintained in these cells. Present in root stem cell daughters and accumulates in maturing root cap layers. Detectable from very early stages of lateral root development. {ECO:0000269|PubMed:19081078, ECO:0000269|PubMed:20197506}. | |||||
| Uniprot | TISSUE SPECIFICITY: Accumulates in maturing root cap cells, in both COL and LRC cells. {ECO:0000269|PubMed:19081078, ECO:0000269|PubMed:20197506}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| TAIR | NAC-domain protein. Involved in root cap development. Involved in a regulatory feedback loop with FEZ. FEZ activates SMB in hte root cap daughter cells soon after division, and SMB in turn represses FEZ expression in these cells, thereby preventing further stem cell divisions. | |||||
| UniProt | Transcription regulator. Together with BRN1 and BRN2, regulates cellular maturation of root cap. Represses stem cell-like divisions in the root cap daughter cells, and thus promotes daughter cell fate. Inhibits expression of its positive regulator FEZ in a feedback loop for controlled switches in cell division plane. Promotes the expression of genes involved in secondary cell walls (SCW) biosynthesis. {ECO:0000269|PubMed:19081078, ECO:0000269|PubMed:20197506}. | |||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
|
||||||
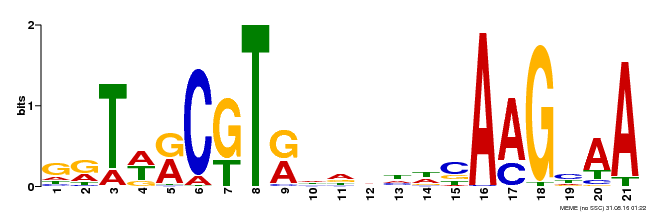
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00250 | DAP | 27203113 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT1G79580.2 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By FEZ in oriented-divised root cap stem cells. {ECO:0000269|PubMed:19081078}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Regulation -- ATRM (Manually Curated Upstream Regulators) ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator (A: Activate/R: Repress) | |||||
| ATRM | AT1G26870 (A) | |||||
| Regulation -- ATRM (Manually Curated Target Genes) ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Target Gene (A: Activate/R: Repress) | |||||
| ATRM | AT1G26870(R) | |||||
| Phenotype -- Disruption Phenotype ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | DISRUPTION PHENOTYPE: Additional root cap layers; increased number of columella (COL) and lateral root cap (LRC) cell layers in the mature embryo and in the postembryonic state. Lateral cap cells continue to divide and fail to detach from the root. {ECO:0000269|PubMed:19081078, ECO:0000269|PubMed:20197506}. | |||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT1G79580 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC010793 | 0.0 | AC010793.3 Genomic sequence for Arabidopsis thaliana BAC F20B17 from chromosome I, complete sequence. | |||
| GenBank | CP002684 | 0.0 | CP002684.1 Arabidopsis thaliana chromosome 1 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001319416.1 | 0.0 | NAC (No Apical Meristem) domain transcriptional regulator superfamily protein | ||||
| Refseq | NP_001323331.1 | 0.0 | NAC (No Apical Meristem) domain transcriptional regulator superfamily protein | ||||
| Refseq | NP_178076.1 | 0.0 | NAC (No Apical Meristem) domain transcriptional regulator superfamily protein | ||||
| Refseq | NP_974178.1 | 0.0 | NAC (No Apical Meristem) domain transcriptional regulator superfamily protein | ||||
| Refseq | NP_974179.1 | 0.0 | NAC (No Apical Meristem) domain transcriptional regulator superfamily protein | ||||
| Swissprot | Q9MA17 | 0.0 | SMB_ARATH; Protein SOMBRERO | ||||
| TrEMBL | A0A178WJ15 | 0.0 | A0A178WJ15_ARATH; URP7 | ||||
| STRING | AT1G79580.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT1G79580.2 |
| Entrez Gene | 844296 |
| iHOP | AT1G79580 |
| wikigenes | AT1G79580 |



