 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT1G75240.1 | ||||||||
| Common Name | AtHB33, F22H5.4, HB33, ZHD5 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | ZF-HD | ||||||||
| Protein Properties | Length: 309aa MW: 33892.8 Da PI: 8.5774 | ||||||||
| Description | homeobox protein 33 | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | ZF-HD_dimer | 103.4 | 1.5e-32 | 72 | 127 | 2 | 58 |
ZF-HD_dimer 2 ekvrYkeClkNhAaslGghavDGCgEfmpsegeegtaaalkCaACgCHRnFHRreve 58
+vrY+eClkNhAas+Gg + DGCgEfmps geegt++al+CaAC+CHRnFHR+e++
AT1G75240.1 72 PTVRYRECLKNHAASVGGSVHDGCGEFMPS-GEEGTIEALRCAACDCHRNFHRKEMD 127
579**************************9.999********************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| ProDom | PD125774 | 4.0E-27 | 73 | 127 | IPR006456 | ZF-HD homeobox protein, Cys/His-rich dimerisation domain |
| Pfam | PF04770 | 1.2E-29 | 74 | 126 | IPR006456 | ZF-HD homeobox protein, Cys/His-rich dimerisation domain |
| TIGRFAMs | TIGR01566 | 2.9E-30 | 75 | 126 | IPR006456 | ZF-HD homeobox protein, Cys/His-rich dimerisation domain |
| PROSITE profile | PS51523 | 26.276 | 76 | 125 | IPR006456 | ZF-HD homeobox protein, Cys/His-rich dimerisation domain |
| Gene3D | G3DSA:1.10.10.60 | 8.3E-31 | 232 | 301 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 1.51E-19 | 234 | 301 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01565 | 2.0E-29 | 240 | 297 | IPR006455 | Homeodomain, ZF-HD class |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009738 | Biological Process | abscisic acid-activated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000013 | anatomy | cauline leaf | ||||
| PO:0000037 | anatomy | shoot apex | ||||
| PO:0000293 | anatomy | guard cell | ||||
| PO:0008019 | anatomy | leaf lamina base | ||||
| PO:0009006 | anatomy | shoot system | ||||
| PO:0009009 | anatomy | plant embryo | ||||
| PO:0009010 | anatomy | seed | ||||
| PO:0009025 | anatomy | vascular leaf | ||||
| PO:0009029 | anatomy | stamen | ||||
| PO:0009030 | anatomy | carpel | ||||
| PO:0009031 | anatomy | sepal | ||||
| PO:0009032 | anatomy | petal | ||||
| PO:0009046 | anatomy | flower | ||||
| PO:0009052 | anatomy | flower pedicel | ||||
| PO:0020038 | anatomy | petiole | ||||
| PO:0020100 | anatomy | hypocotyl | ||||
| PO:0025022 | anatomy | collective leaf structure | ||||
| PO:0001054 | developmental stage | vascular leaf senescent stage | ||||
| PO:0001078 | developmental stage | plant embryo cotyledonary stage | ||||
| PO:0001081 | developmental stage | mature plant embryo stage | ||||
| PO:0001185 | developmental stage | plant embryo globular stage | ||||
| PO:0004507 | developmental stage | plant embryo bilateral stage | ||||
| PO:0007064 | developmental stage | LP.12 twelve leaves visible stage | ||||
| PO:0007115 | developmental stage | LP.04 four leaves visible stage | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| PO:0007616 | developmental stage | flowering stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 309 aa Download sequence Send to blast |
MDMRSHEMIE RRREDNGNNN GGVVISNIIS TNIDDNCNGN NNNTRVSCNS QTLDHHQSKS 60 PSSFSISAAA KPTVRYRECL KNHAASVGGS VHDGCGEFMP SGEEGTIEAL RCAACDCHRN 120 FHRKEMDGVG SSDLISHHRH HHYHHNQYGG GGGRRPPPPN MMLNPLMLPP PPNYQPIHHH 180 KYGMSPPGGG GMVTPMSVAY GGGGGGAESS SEDLNLYGQS SGEGAGAAAG QMAFSMSSSK 240 KRFRTKFTTD QKERMMDFAE KLGWRMNKQD EEELKRFCGE IGVKRQVFKV WMHNNKNNAK 300 KPPTPTTTL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wh7_A | 9e-23 | 239 | 299 | 16 | 76 | ZF-HD homeobox family protein |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| At.34802 | 0.0 | flower| leaf| seed | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 145337570 | 0.0 | ||||
| Genevisible | 256452_at | 0.0 | ||||
| Expression Atlas | AT1G75240 | - | ||||
| AtGenExpress | AT1G75240 | - | ||||
| ATTED-II | AT1G75240 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Mostly expressed in flowers and inflorescence. {ECO:0000269|PubMed:16428600}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Putative transcription factor. Binds DNA at 5'-ATTA-3' consensus promoter regions. Regulates floral architecture and leaf development. Regulators in the abscisic acid (ABA) signal pathway that confers sensitivity to ABA in an ARF2-dependent manner. {ECO:0000269|PubMed:16428600, ECO:0000269|PubMed:21059647, ECO:0000269|PubMed:21779177}. | |||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
|
||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00238 | DAP | 27203113 | Download |
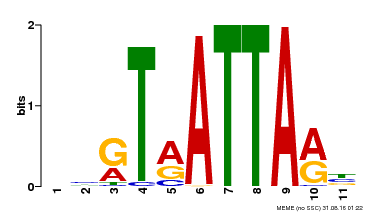
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT1G75240.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Repressed by abscisic acid (ABA) and ARF2. {ECO:0000269|PubMed:21779177}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Interaction ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Intact With | |||||
| IntAct | Search Q9FRL5 | |||||
| Phenotype -- Disruption Phenotype ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | DISRUPTION PHENOTYPE: Increased resistance to abscisic acid (ABA) in the seed germination and primary root growth. {ECO:0000269|PubMed:21779177}. | |||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT1G75240 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC025814 | 0.0 | AC025814.7 Arabidopsis thaliana chromosome 1 BAC F22H5 genomic sequence, complete sequence. | |||
| GenBank | AK229038 | 0.0 | AK229038.1 Arabidopsis thaliana mRNA for hypothetical protein, complete cds, clone: RAFL16-33-K14. | |||
| GenBank | AY084462 | 0.0 | AY084462.1 Arabidopsis thaliana clone 10873 mRNA, complete sequence. | |||
| GenBank | BT028894 | 0.0 | BT028894.1 Arabidopsis thaliana unknown protein (At1g75240) mRNA, complete cds. | |||
| GenBank | CP002684 | 0.0 | CP002684.1 Arabidopsis thaliana chromosome 1 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_565106.1 | 0.0 | homeobox protein 33 | ||||
| Swissprot | Q9FRL5 | 0.0 | ZHD5_ARATH; Zinc-finger homeodomain protein 5 | ||||
| TrEMBL | A0A178WFZ4 | 0.0 | A0A178WFZ4_ARATH; ZHD5 | ||||
| STRING | AT1G75240.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM7279 | 25 | 43 | Representative plant | OGRP91 | 16 | 237 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT1G75240.1 |
| Entrez Gene | 843861 |
| iHOP | AT1G75240 |
| wikigenes | AT1G75240 |



