| Signature Domain? help Back to Top |
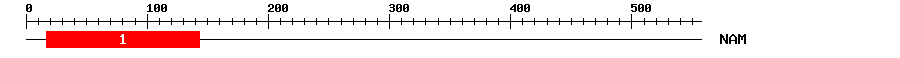 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | NAM | 158.7 | 2.3e-49 | 17 | 143 | 2 | 128 |
NAM 2 ppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLp.k.kvkaeekewyfFskrdkkyatgkrknratksgyWkatgkdkevlskkg 97
+pGfrFhPtdeelv++yLk+k+ +k+l++ +vi vd+yk++P++Lp + ++k+++++w++F++r++ky++ r+nr t++gyWkatgkd+ + ++
AT1G34190.1 17 APGFRFHPTDEELVMYYLKRKICRKRLRV-NVIGVVDVYKMDPEELPgQsMLKTGDRQWFYFTPRSRKYPNAARSNRGTENGYWKATGKDRVIEY-NS 112
69***************************.99**************96347888999************************************99.99 PP
NAM 98 elvglkktLvfykgrapkgektdWvmheyrl 128
+ vglkktLvfy+grap+ge+tdWvmhey++
AT1G34190.1 113 RSVGLKKTLVFYRGRAPSGERTDWVMHEYTM 143
9****************************97 PP
|
| Publications
? help Back to Top |
- Riechmann JL, et al.
Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes.
Science, 2000. 290(5499): p. 2105-10
[PMID:11118137] - Yamada K, et al.
Empirical analysis of transcriptional activity in the Arabidopsis genome.
Science, 2003. 302(5646): p. 842-6
[PMID:14593172] - Hu W,Wang Y,Bowers C,Ma H
Isolation, sequence analysis, and expression studies of florally expressed cDNAs in Arabidopsis.
Plant Mol. Biol., 2003. 53(4): p. 545-63
[PMID:15010618] - Ooka H, et al.
Comprehensive analysis of NAC family genes in Oryza sativa and Arabidopsis thaliana.
DNA Res., 2003. 10(6): p. 239-47
[PMID:15029955] - Duarte JM, et al.
Expression pattern shifts following duplication indicative of subfunctionalization and neofunctionalization in regulatory genes of Arabidopsis.
Mol. Biol. Evol., 2006. 23(2): p. 469-78
[PMID:16280546] - Kim SY, et al.
Exploring membrane-associated NAC transcription factors in Arabidopsis: implications for membrane biology in genome regulation.
Nucleic Acids Res., 2007. 35(1): p. 203-13
[PMID:17158162] - Barakat A,Wall PK,Diloreto S,Depamphilis CW,Carlson JE
Conservation and divergence of microRNAs in Populus.
BMC Genomics, 2007. 8: p. 481
[PMID:18166134] - Huang YC,Chang YL,Hsu JJ,Chuang HW
Transcriptome analysis of auxin-regulated genes of Arabidopsis thaliana.
Gene, 2008. 420(2): p. 118-24
[PMID:18577427] - Ascencio-Ib
Global analysis of Arabidopsis gene expression uncovers a complex array of changes impacting pathogen response and cell cycle during geminivirus infection.
Plant Physiol., 2008. 148(1): p. 436-54
[PMID:18650403] - Wang Y, et al.
Transcriptome analyses show changes in gene expression to accompany pollen germination and tube growth in Arabidopsis.
Plant Physiol., 2008. 148(3): p. 1201-11
[PMID:18775970] - Ng S, et al.
A membrane-bound NAC transcription factor, ANAC017, mediates mitochondrial retrograde signaling in Arabidopsis.
Plant Cell, 2013. 25(9): p. 3450-71
[PMID:24045017] - Hofmann NR
Endoplasmic reticulum-localized transcription factors and mitochondrial retrograde regulation.
Plant Cell, 2013. 25(9): p. 3151
[PMID:24045018] - Van Aken O, et al.
Mitochondrial and Chloroplast Stress Responses Are Modulated in Distinct Touch and Chemical Inhibition Phases.
Plant Physiol., 2016. 171(3): p. 2150-65
[PMID:27208304] - Van Aken O,Ford E,Lister R,Huang S,Millar AH
Retrograde signalling caused by heritable mitochondrial dysfunction is partially mediated by ANAC017 and improves plant performance.
Plant J., 2016. 88(4): p. 542-558
[PMID:27425258] - Hu Z, et al.
Mitochondrial Defects Confer Tolerance against Cellulose Deficiency.
Plant Cell, 2016. 28(9): p. 2276-2290
[PMID:27543091] - Chi YH, et al.
The membrane-tethered NAC transcription factor, AtNTL7, contributes to ER-stress resistance in Arabidopsis.
Biochem. Biophys. Res. Commun., 2017. 488(4): p. 641-647
[PMID:28088515] - Van Aken O,Pogson BJ
Convergence of mitochondrial and chloroplastic ANAC017/PAP-dependent retrograde signalling pathways and suppression of programmed cell death.
Cell Death Differ., 2017. 24(6): p. 955-960
[PMID:28498364] - Cheng P, et al.
The ERA-Related GTPase AtERG2 Associated with Mitochondria 18S RNA Is Essential for Early Embryo Development in Arabidopsis.
Front Plant Sci, 2018. 9: p. 182
[PMID:29497438]
|