 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT1G32510.1 | ||||||||
| Common Name | ANAC011, F5D14.30, NAC011 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 283aa MW: 32871.8 Da PI: 5.1094 | ||||||||
| Description | NAC domain containing protein 11 | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 161.3 | 3.7e-50 | 6 | 140 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLp.k.kvkaeekewyfFskrdkkyatgkrknratksgyWkatgkdkevlsk. 95
lppGfrF Ptdeelv +yL+++ eg ++el e+i+ +d+yk++Pw+Lp k + +++ ew+fF++rd+ky++g+r nratksgyWkatgkd++++++
AT1G32510.1 6 LPPGFRFYPTDEELVGYYLHRRNEGLEIEL-EIIPLMDLYKFDPWELPeKsFLPNRDMEWFFFCHRDRKYQNGSRINRATKSGYWKATGKDRKIVCHs 102
79****************************.99**************95334556888*************************************986 PP
NAM 96 .....kgelvglkktLvfykgrapkgektdWvmheyrl 128
+++ +g++ktLvfy grap g +t+Wvmheyrl
AT1G32510.1 103 sssssSSSITGCRKTLVFYMGRAPFGGRTEWVMHEYRL 140
666555555***************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 6.67E-57 | 4 | 164 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 55.318 | 6 | 164 | IPR003441 | NAC domain |
| Pfam | PF02365 | 1.8E-27 | 7 | 140 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 283 aa Download sequence Send to blast |
MVGSFLPPGF RFYPTDEELV GYYLHRRNEG LEIELEIIPL MDLYKFDPWE LPEKSFLPNR 60 DMEWFFFCHR DRKYQNGSRI NRATKSGYWK ATGKDRKIVC HSSSSSSSSS ITGCRKTLVF 120 YMGRAPFGGR TEWVMHEYRL FDNDTSQGSL NFKGDFALCR VIKRNEHTLK KCEIISPEVS 180 DESLSNNVNN FCQASDLEKG SCDASNTRLS SPDFILESSF QGNSHSKTEE DSGFQVFTLP 240 EFEYPLEVFA DLNFDLEMED PFMFDYHPEP HMNNEVMSHH IRG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ut4_A | 8e-51 | 6 | 170 | 17 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut4_B | 8e-51 | 6 | 170 | 17 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut7_A | 8e-51 | 6 | 170 | 17 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut7_B | 8e-51 | 6 | 170 | 17 | 171 | NO APICAL MERISTEM PROTEIN |
| 3swm_A | 8e-51 | 6 | 170 | 20 | 174 | NAC domain-containing protein 19 |
| 3swm_B | 8e-51 | 6 | 170 | 20 | 174 | NAC domain-containing protein 19 |
| 3swm_C | 8e-51 | 6 | 170 | 20 | 174 | NAC domain-containing protein 19 |
| 3swm_D | 8e-51 | 6 | 170 | 20 | 174 | NAC domain-containing protein 19 |
| 3swp_A | 8e-51 | 6 | 170 | 20 | 174 | NAC domain-containing protein 19 |
| 3swp_B | 8e-51 | 6 | 170 | 20 | 174 | NAC domain-containing protein 19 |
| 3swp_C | 8e-51 | 6 | 170 | 20 | 174 | NAC domain-containing protein 19 |
| 3swp_D | 8e-51 | 6 | 170 | 20 | 174 | NAC domain-containing protein 19 |
| 4dul_A | 8e-51 | 6 | 170 | 17 | 171 | NAC domain-containing protein 19 |
| 4dul_B | 8e-51 | 6 | 170 | 17 | 171 | NAC domain-containing protein 19 |
| Search in ModeBase | ||||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| Expression Atlas | AT1G32510 | |||||
| AtGenExpress | AT1G32510 | |||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, rosettes leaves, cauline leaves and stems. {ECO:0000269|PubMed:24285786}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator involved in the positive regulation of abscisic acid (ABA) responsive genes. Acts as a positive factor of ABA-mediated responses. Involved in the transcriptional activation of ABA-inducible genes in response to dehydration and osmotic stresses. Plays a positive role in both stomatal closure and water loss under dehydration stress conditions. Acts synergistically with ABF2 to activate the dehydration stress-response factor RD29A transcription. Binds to the consensus core cis-acting elements 5'-CGTA-3' and 5'-CACG-3' at the RD29A promoter (PubMed:24285786). Involved in hypocotyl graft union formation. Required for the auxin-mediated promotion of vascular tissue proliferation during hypocotyl graft attachment (PubMed:25182467). {ECO:0000269|PubMed:24285786, ECO:0000269|PubMed:25182467}. | |||||
| UniProt | Transcription factor involved in tissue reunion of wounded inflorescence stems. Required for the division of pith cells in the reunion process, which is dependent on polar-transported auxin and the wound-inducible hormones ethylene and jasmonate (PubMed:21911380). Binds to the promoters of XTH19 and XTH20 to induce their expression via auxin signaling. XTH19 and XTH20 are involved in cell proliferation in the tissue reunion process of incised stems (PubMed:25182467). Involved in hypocotyl graft union formation. Required for the auxin- mediated promotion of vascular tissue proliferation during hypocotyl graft attachment (PubMed:27986917). {ECO:0000269|PubMed:21911380, ECO:0000269|PubMed:25182467, ECO:0000269|PubMed:27986917}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00175 | DAP | 27203113 | Download |
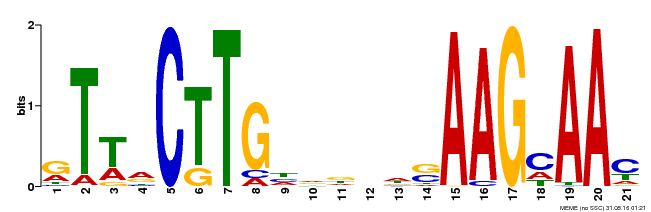
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT1G32510.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by abscisic acid (ABA), dehydration and osmotic stress. {ECO:0000269|PubMed:24285786}. | |||||
| UniProt | INDUCTION: Induced by wounding in the flowering stem. {ECO:0000269|PubMed:21911380}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT1G32510 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | CP002684 | 1e-170 | CP002684.1 Arabidopsis thaliana chromosome 1 sequence. | |||
| GenBank | F5D14 | 1e-170 | AC007767.3 Sequence of BAC F5D14 from Arabidopsis thaliana chromosome 1, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_174529.2 | 0.0 | NAC domain containing protein 11 | ||||
| Swissprot | O49697 | 8e-86 | NAC71_ARATH; NAC domain-containing protein 71 | ||||
| Swissprot | Q9LS24 | 2e-85 | NAC96_ARATH; NAC domain-containing protein 96 | ||||
| TrEMBL | A0A178WL27 | 0.0 | A0A178WL27_ARATH; NAC011 | ||||
| TrEMBL | Q9LQK5 | 0.0 | Q9LQK5_ARATH; F5D14.30 protein | ||||
| STRING | AT1G32510.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM15029 | 17 | 19 | Representative plant | OGRP17 | 15 | 800 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT1G32510.1 |
| Entrez Gene | 840145 |
| iHOP | AT1G32510 |
| wikigenes | AT1G32510 |



