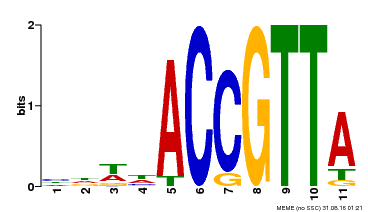
| Signature Domain? help Back to Top |
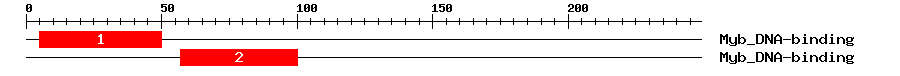 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | Myb_DNA-binding | 56.3 | 7.3e-18 | 5 | 50 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
rg W + Ede+l ++v+q+G+++W++Ia+++ gR++k+c++rw++
AT1G17950.1 5 RGHWRPAEDEKLRELVEQFGPHNWNAIAQKLS-GRSGKSCRLRWFNQ 50
899*****************************.***********996 PP
|
| 2 | Myb_DNA-binding | 55.5 | 1.3e-17 | 57 | 100 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
r+++T+eE+e+l+ ++ +G++ W+ Iar ++ gRt++ +k++w+
AT1G17950.1 57 RNPFTEEEEERLLASHRIHGNR-WSVIARFFP-GRTDNAVKNHWHV 100
789*******************.*********.***********96 PP
|
| Publications
? help Back to Top |
- Riechmann JL, et al.
Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes.
Science, 2000. 290(5499): p. 2105-10
[PMID:11118137] - Stracke R,Werber M,Weisshaar B
The R2R3-MYB gene family in Arabidopsis thaliana.
Curr. Opin. Plant Biol., 2001. 4(5): p. 447-56
[PMID:11597504] - Czechowski T,Bari RP,Stitt M,Scheible WR,Udvardi MK
Real-time RT-PCR profiling of over 1400 Arabidopsis transcription factors: unprecedented sensitivity reveals novel root- and shoot-specific genes.
Plant J., 2004. 38(2): p. 366-79
[PMID:15078338] - Zhao C,Craig JC,Petzold HE,Dickerman AW,Beers EP
The xylem and phloem transcriptomes from secondary tissues of the Arabidopsis root-hypocotyl.
Plant Physiol., 2005. 138(2): p. 803-18
[PMID:15923329] - Duarte JM, et al.
Expression pattern shifts following duplication indicative of subfunctionalization and neofunctionalization in regulatory genes of Arabidopsis.
Mol. Biol. Evol., 2006. 23(2): p. 469-78
[PMID:16280546] - Andersson-Gunner
Biosynthesis of cellulose-enriched tension wood in Populus: global analysis of transcripts and metabolites identifies biochemical and developmental regulators in secondary wall biosynthesis.
Plant J., 2006. 45(2): p. 144-65
[PMID:16367961] - Yanhui C, et al.
The MYB transcription factor superfamily of Arabidopsis: expression analysis and phylogenetic comparison with the rice MYB family.
Plant Mol. Biol., 2006. 60(1): p. 107-24
[PMID:16463103] - Cao D,Cheng H,Wu W,Soo HM,Peng J
Gibberellin mobilizes distinct DELLA-dependent transcriptomes to regulate seed germination and floral development in Arabidopsis.
Plant Physiol., 2006. 142(2): p. 509-25
[PMID:16920880] - Ko JH,Beers EP,Han KH
Global comparative transcriptome analysis identifies gene network regulating secondary xylem development in Arabidopsis thaliana.
Mol. Genet. Genomics, 2006. 276(6): p. 517-31
[PMID:16969662] - Ko JH,Yang SH,Park AH,Lerouxel O,Han KH
ANAC012, a member of the plant-specific NAC transcription factor family, negatively regulates xylary fiber development in Arabidopsis thaliana.
Plant J., 2007. 50(6): p. 1035-48
[PMID:17565617] - Zhong R,Lee C,Zhou J,McCarthy RL,Ye ZH
A battery of transcription factors involved in the regulation of secondary cell wall biosynthesis in Arabidopsis.
Plant Cell, 2008. 20(10): p. 2763-82
[PMID:18952777] - Cheng H, et al.
Gibberellin acts through jasmonate to control the expression of MYB21, MYB24, and MYB57 to promote stamen filament growth in Arabidopsis.
PLoS Genet., 2009. 5(3): p. e1000440
[PMID:19325888] - Ko JH,Kim WC,Han KH
Ectopic expression of MYB46 identifies transcriptional regulatory genes involved in secondary wall biosynthesis in Arabidopsis.
Plant J., 2009. 60(4): p. 649-65
[PMID:19674407] - Park MY,Kang JY,Kim SY
Overexpression of AtMYB52 confers ABA hypersensitivity and drought tolerance.
Mol. Cells, 2011. 31(5): p. 447-54
[PMID:21399993] - Zhong R,Ye ZH
MYB46 and MYB83 bind to the SMRE sites and directly activate a suite of transcription factors and secondary wall biosynthetic genes.
Plant Cell Physiol., 2012. 53(2): p. 368-80
[PMID:22197883] - Meinke DW
A survey of dominant mutations in Arabidopsis thaliana.
Trends Plant Sci., 2013. 18(2): p. 84-91
[PMID:22995285] - Cassan-Wang H, et al.
Identification of novel transcription factors regulating secondary cell wall formation in Arabidopsis.
Front Plant Sci, 2013. 4: p. 189
[PMID:23781226] - Jin J, et al.
An Arabidopsis Transcriptional Regulatory Map Reveals Distinct Functional and Evolutionary Features of Novel Transcription Factors.
Mol. Biol. Evol., 2015. 32(7): p. 1767-73
[PMID:25750178] - Shi D, et al.
MYB52 Negatively Regulates Pectin Demethylesterification in Seed Coat Mucilage.
Plant Physiol., 2018. 176(4): p. 2737-2749
[PMID:29440562] - Kranz HD, et al.
Towards functional characterisation of the members of the R2R3-MYB gene family from Arabidopsis thaliana.
Plant J., 1998. 16(2): p. 263-76
[PMID:9839469]
|





