 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | EMT25190 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Aegilops
|
||||||||
| Family | Nin-like | ||||||||
| Protein Properties | Length: 944aa MW: 103467 Da PI: 6.0724 | ||||||||
| Description | Nin-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | RWP-RK | 82.3 | 4.9e-26 | 583 | 645 | 1 | 52 |
RWP-RK 1 aekeisledlskyFslpikdAAkeLg...........vclTvLKriCRqyGIkRWPhRkiksl 52
aek+isle+l++yFs+++k+AAk+Lg +c+T++KriCRq+GI+RWP+Rki+++
EMT25190 583 AEKTISLEVLQQYFSGSLKNAAKSLGglkwqqnvaacMCPTTMKRICRQHGISRWPSRKINKV 645
589**********************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51519 | 14.845 | 573 | 665 | IPR003035 | RWP-RK domain |
| Pfam | PF02042 | 2.5E-23 | 586 | 645 | IPR003035 | RWP-RK domain |
| SuperFamily | SSF54277 | 5.23E-22 | 841 | 934 | No hit | No description |
| PROSITE profile | PS51745 | 19.189 | 850 | 932 | IPR000270 | PB1 domain |
| Gene3D | G3DSA:3.10.20.240 | 1.2E-25 | 850 | 931 | No hit | No description |
| SMART | SM00666 | 2.2E-24 | 850 | 932 | IPR000270 | PB1 domain |
| Pfam | PF00564 | 5.9E-15 | 850 | 931 | IPR000270 | PB1 domain |
| CDD | cd06407 | 1.03E-33 | 851 | 931 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0010167 | Biological Process | response to nitrate | ||||
| GO:0042128 | Biological Process | nitrate assimilation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 944 aa Download sequence Send to blast |
MDIDPSSVSH ADGGGDLWPF DSLTTSLFFS SVSSSPPLHP LPVASSSWLT PPSPLWLFED 60 RQMMPIEVGP APAATPDNTA AVAEEAQRAR SGNSDTPVKK TERLTNKWQF NLALHDDSTN 120 SSCLFKEKLT HALRYFKEST DQHLLVQVWA PVKSGDRYVL TTSGQPFVLD HQSIGLLQYR 180 AVSMMYMFSV DGDNAGELGL PGRVYKQKVP EWTPNVQYYS STEYPRLNHA ISYNVHGTVA 240 LPVFDPSVQS CIAVVELIMT SKKINYADEV DKVCKALEAV NLKSTEILEH PNVQICNEGR 300 QSALVEILEI LTVVCEEHKL PLAQTWVPCK YRSVLAHGGG VKKSCLSFDG SCMGEVCMST 360 SDVAFHVIDA HMWGFRDACV EHHLQKGQGV SGKAFIYHRP CFSKDISQFC KLEYPLVHYA 420 RMFGLAGCFA ICLQSPYTGD DYYMLEFFLP PSCKEEDDQN ALLESILGLI NQCLRNLKVA 480 GNGESNEASL QLSNVIMIES EDFKTNVHFE NSEGFRESPE GDTHGGAHEF DKANSKVSEG 540 HLLADDNSQN NGVSVSRPNG SAASDSSLLH KNGKPPERRR GKAEKTISLE VLQQYFSGSL 600 KNAAKSLGGL KWQQNVAACM CPTTMKRICR QHGISRWPSR KINKVNRSLS KLKQVIESVQ 660 GSDAAFNLTS ITGPLPTIPV GPSSDSFNKE KASESKAEHS NRAVDGDRDS SLQKSQENGS 720 HFGALMSQQG FADTGNNVQL EADKASLSRS SSGEGSINSR TSEGSCQGSP ANQTFVCQPI 780 ASMFLEPQEP QLNPEGFTKE PFQEPELPLS RMLIEDSGSS KDLKNLFGSA IGQPMLAPPS 840 NFGPMRNSGT VTIKASFKED IVRFRFPCSS SVMALKDEVA KRLRMDAGMF DIKYLDDDHE 900 WVKLACNADL EECIEISRHS GTHVIRLLVS DVAAHIGSSC GSSG |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
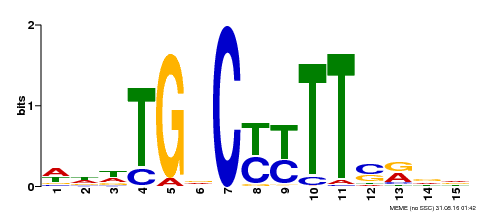
| Motif ID | Method | Source | Motif file |
| MP00449 | DAP | Transfer from AT4G24020 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HG670306 | 0.0 | HG670306.1 Triticum aestivum chromosome 3B, genomic scaffold, cultivar Chinese Spring. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020163903.1 | 0.0 | protein NLP3-like | ||||
| Swissprot | Q5NB82 | 0.0 | NLP3_ORYSJ; Protein NLP3 | ||||
| TrEMBL | M8BJK9 | 0.0 | M8BJK9_AEGTA; Uncharacterized protein | ||||
| STRING | EMT25190 | 0.0 | (Aegilops tauschii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP4778 | 36 | 60 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G64530.1 | 0.0 | Nin-like family protein | ||||



