 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | EMT22828 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Aegilops
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 402aa MW: 43148.8 Da PI: 6.0588 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 102.1 | 3.5e-32 | 230 | 285 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
kpr++W+peLH+rF++a++qLGGs++AtPk+i+elmkv+gLt+++vkSHLQkYRl+
EMT22828 230 KPRRCWAPELHRRFLQALQQLGGSHVATPKQIRELMKVDGLTNDEVKSHLQKYRLH 285
79****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 12.822 | 227 | 287 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.1E-28 | 228 | 288 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 7.17E-17 | 228 | 287 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 3.3E-27 | 230 | 285 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.8E-6 | 232 | 283 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 402 aa Download sequence Send to blast |
MEMEMEVPPA QADCDGARRR LQDYLLALEE ERRKIHVFQR ELPLCLDLVT QTIEGMKSQM 60 GGAASEETVS DHGGPVLEEF IPLKPSLSLS SSDDETSNHH AAAPVSNAKA ETPPPTPETK 120 KAMPDWLQSA QLSTWSEPQQ SSSLQKVLPC RPVALNATRT GGAFHPFEKE KQPEPELPAS 180 STTAPASSAL VGDSGDKATS DTEVHDKDSK DVTEKDTDKD KEGHSQPNRK PRRCWAPELH 240 RRFLQALQQL GGSHVATPKQ IRELMKVDGL TNDEVKSHLQ KYRLHTSRRP SSTAQSGATA 300 ATQPAPQFFV VGSCIWVPQQ EYAAAAAAAA AAADATNVAP GSANQVYART ATLPSGLKPH 360 LEKQSSRQSE GPRSGDTGGV NSDTDNPAMS SSSSQTTSAG HP |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 4e-13 | 230 | 283 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 4e-13 | 230 | 283 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 4e-13 | 230 | 283 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 4e-13 | 230 | 283 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 4e-13 | 230 | 283 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 4e-13 | 230 | 283 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 4e-13 | 230 | 283 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 4e-13 | 230 | 283 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional repressor that may play a role in response to nitrogen. May be involved in a time-dependent signaling for transcriptional regulation of nitrate-responsive genes. Binds specifically to the DNA sequence motif 5'-GAATC-3' or 5'-GAATATTC-3'. Represses the activity of its own promoter trough binding to these motifs. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00160 | DAP | Transfer from AT1G25550 | Download |
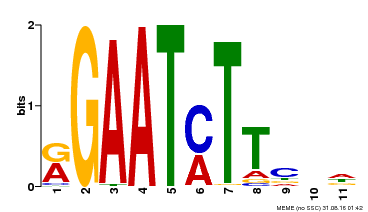
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by nitrate. {ECO:0000269|PubMed:23324170}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK361638 | 0.0 | AK361638.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv1145I13. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020198678.1 | 0.0 | myb family transcription factor EFM-like | ||||
| Refseq | XP_020198679.1 | 0.0 | myb family transcription factor EFM-like | ||||
| Swissprot | Q6Z869 | 1e-128 | NIGT1_ORYSJ; Transcription factor NIGT1 | ||||
| TrEMBL | A0A453AR00 | 0.0 | A0A453AR00_AEGTS; Uncharacterized protein | ||||
| TrEMBL | M8BLZ2 | 0.0 | M8BLZ2_AEGTA; Two-component response regulator ARR2 | ||||
| STRING | EMT22828 | 0.0 | (Aegilops tauschii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6470 | 37 | 54 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G25550.1 | 3e-59 | G2-like family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



