 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | EMT18702 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Aegilops
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 317aa MW: 34403.8 Da PI: 4.4897 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 33.5 | 8.9e-11 | 120 | 171 | 4 | 55 |
XCHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 4 lkrerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkke 55
+k+ rr ++NR +A +sR+RKk +++ Le+k+k Leae ++L l+ e
EMT18702 120 SKKKRRQMRNRDSAMKSRERKKSYVKDLETKSKYLEAECRRLSYALQCCAAE 171
699************************************9998777665555 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 4.4E-6 | 116 | 181 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 9.737 | 119 | 161 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF00170 | 1.2E-9 | 121 | 177 | IPR004827 | Basic-leucine zipper domain |
| Gene3D | G3DSA:1.20.5.170 | 4.9E-12 | 121 | 180 | No hit | No description |
| SuperFamily | SSF57959 | 1.48E-10 | 121 | 179 | No hit | No description |
| CDD | cd14704 | 8.47E-14 | 122 | 173 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 124 | 139 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000165 | Biological Process | MAPK cascade | ||||
| GO:0002679 | Biological Process | respiratory burst involved in defense response | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0006612 | Biological Process | protein targeting to membrane | ||||
| GO:0006984 | Biological Process | ER-nucleus signaling pathway | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0009410 | Biological Process | response to xenobiotic stimulus | ||||
| GO:0009693 | Biological Process | ethylene biosynthetic process | ||||
| GO:0009738 | Biological Process | abscisic acid-activated signaling pathway | ||||
| GO:0009867 | Biological Process | jasmonic acid mediated signaling pathway | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0010286 | Biological Process | heat acclimation | ||||
| GO:0010363 | Biological Process | regulation of plant-type hypersensitive response | ||||
| GO:0030968 | Biological Process | endoplasmic reticulum unfolded protein response | ||||
| GO:0031348 | Biological Process | negative regulation of defense response | ||||
| GO:0043069 | Biological Process | negative regulation of programmed cell death | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005789 | Cellular Component | endoplasmic reticulum membrane | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 317 aa Download sequence Send to blast |
MDTDLDLDAL LASFAGESAA VSELLAPPPL DAAEAGGPEA EEEAAGAEPV DGISVDEFLD 60 TLFDGAEEGG EKGNGSEAEA GGSTDGDSRR GEDGVEVVTP ETEAEVVTPE TEVDGDDPIS 120 KKKRRQMRNR DSAMKSRERK KSYVKDLETK SKYLEAECRR LSYALQCCAA ENMALRQNML 180 KDRPIGAHTA MQESAVLSEL SELDVSELKS LHEMQPGLLA RRNLSCAKCN CFHFAEPLPL 240 VSLLWLVSIV CLFLTPGLPN RSLVAPRRAE RDLAMVAGKP SSDQPETLEL LLHGRRWRGT 300 RERIKLDTPP LRAAAAC |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 120 | 124 | KKKRR |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in endoplasmic reticulum (ER) stress response (PubMed:21223397, PubMed:22050533, PubMed:22199238). Acts downstream of the ER stress sensors IRE1, BZIP39 and BZIP60 to activate BiP chaperone genes (PubMed:22050533, PubMed:22199238). {ECO:0000269|PubMed:21223397, ECO:0000269|PubMed:22050533, ECO:0000269|PubMed:22199238}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00019 | PBM | Transfer from AT1G42990 | Download |
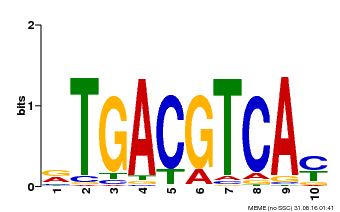
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By dithiothreitol-induced endoplasmic reticulum (ER) stress response (PubMed:22050533, PubMed:21223397). Induced by salt stress (PubMed:22050533). {ECO:0000269|PubMed:21223397, ECO:0000269|PubMed:22050533}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK365505 | 1e-136 | AK365505.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv2034K24. | |||
| GenBank | AK366328 | 1e-136 | AK366328.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv2041P19. | |||
| GenBank | AK369957 | 1e-136 | AK369957.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv2101P04. | |||
| GenBank | AK370149 | 1e-136 | AK370149.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv2105L10. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020184631.1 | 1e-179 | bZIP transcription factor 50 | ||||
| Swissprot | Q69XV0 | 2e-87 | BZP50_ORYSJ; bZIP transcription factor 50 | ||||
| TrEMBL | R7W983 | 0.0 | R7W983_AEGTA; BZIP transcription factor 60 | ||||
| STRING | EMT18702 | 0.0 | (Aegilops tauschii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6934 | 37 | 50 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G42990.1 | 4e-11 | basic region/leucine zipper motif 60 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



