 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | EMT16107 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Aegilops
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1030aa MW: 114613 Da PI: 6.2268 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 178.8 | 6.5e-56 | 21 | 136 | 3 | 117 |
CG-1 3 ke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcywlL 102
+e ++rwl+++ei++iL n++k++++ e+++rp+sgsl+L++rk++ryfrkDG+ w+kkkdgktv+E+hekLKvg+v+vl+cyYah+een++fqrr+ywlL
EMT16107 21 QEaQNRWLRPTEICQILSNYKKFSIAPEPPNRPPSGSLFLFDRKILRYFRKDGHIWRKKKDGKTVKEAHEKLKVGSVDVLHCYYAHGEENENFQRRTYWLL 121
5559************************************************************************************************* PP
CG-1 103 eeelekivlvhylev 117
ee + +ivlvhyle
EMT16107 122 EEGFMNIVLVHYLEI 136
************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 81.103 | 16 | 142 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 3.2E-77 | 19 | 137 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 2.2E-50 | 22 | 135 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 1.6E-7 | 440 | 527 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 2.8E-7 | 441 | 514 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 5.44E-18 | 441 | 526 | IPR014756 | Immunoglobulin E-set |
| SuperFamily | SSF48403 | 1.51E-16 | 618 | 733 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.61E-12 | 619 | 731 | No hit | No description |
| Gene3D | G3DSA:1.25.40.20 | 5.9E-17 | 628 | 734 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 17.396 | 639 | 733 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 3.3E-6 | 645 | 734 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 0.069 | 672 | 701 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 9.458 | 672 | 704 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 420 | 711 | 740 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 3.72E-8 | 843 | 896 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 1.5 | 845 | 867 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.627 | 847 | 875 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0055 | 847 | 866 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0022 | 868 | 890 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.67 | 869 | 893 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 6.5E-5 | 870 | 890 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1030 aa Download sequence Send to blast |
MAEMHKYGLS NQPPDIPQIL QEAQNRWLRP TEICQILSNY KKFSIAPEPP NRPPSGSLFL 60 FDRKILRYFR KDGHIWRKKK DGKTVKEAHE KLKVGSVDVL HCYYAHGEEN ENFQRRTYWL 120 LEEGFMNIVL VHYLEIKGGK QSFSRSKEAE DSPACSNSFA SQSQVASQTM DAESPYSGQI 180 SEYEDAETGY QDENQARTAN ISNNFVTHHG IASVFNDVGA GLRSGSKTAL DSVHFGEPFP 240 EYPTGFTEPT LYSSVTTMGS NNLDDNSRLE TLMTEALYTN NLTQKETDAL SAAGMTSSQR 300 RPPEAPVHSG RLKRADIVKR GLKDWNITKE LAMDRGAWKL AIHVHNDSYT DGSMGYPLLK 360 QSSLDLFKIE PNGLKKFDSF SKWMSDELAA DLDIKSTSDA FWSSTETVNV ADGSSMSIPM 420 NEQLDAYVVS PSLSQDQLFS IIDVSPSWAY TGSQNKVLIT GTFLTNKEHV ENCKWSCMFG 480 DVEVPVEVLA DGSLRCYTPV HQSGRVPFYV TCSNRVACSE VREFEFHDSE TQHMEAADPH 540 ITGINEMHLH IRLEKLLSLG PDDYEKYVMS GGNEKSELIS TIGSLMLDDK FTNLSAPSDE 600 EFSAAQDKNL EKSVKDKLYY WLIHKIHDDG KGPNVLGKEG QGVIHLVAAL GYDWAIRPII 660 AAGVHVNFRD VRGWTALHWA ASCGRERTVG ALITNGAAAG ALTDPTPHFL SGRTPADLAS 720 DNGHKGIAGF LAESALTSHL SALTLKEAKG CNVEEICGSA EADGFAEPTS AQLSRQDSQA 780 ESLKDSLSAV RKSTLAASKI FQAFRVESFH RKKVVEYGDD DCGLSDERTL SLVSLKNTKS 840 GQNDMPHSAA VRIQNKFRGW KGRKEFMIIR QKIIKIQAHI RGHQVRRNYK KVIWSVGIVE 900 KVILRWRRKG RGLRGFQPDK QLEGPSSQIQ PAEGGSAEAE DEYDFLKDGR KQAEGRLQRS 960 LARVKSMTQY PEAREQYSRL QACVTELQES KAIQDKMLSD AAGGDGGDFM VDLEDLCADE 1020 LLDTPMSTIL |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
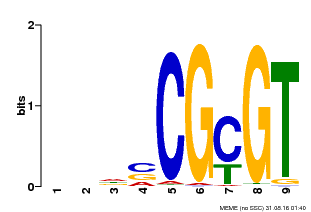
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020196708.1 | 0.0 | calmodulin-binding transcription activator 2-like isoform X1 | ||||
| TrEMBL | N1QYH5 | 0.0 | N1QYH5_AEGTA; Calmodulin-binding transcription activator 2 | ||||
| STRING | EMT16107 | 0.0 | (Aegilops tauschii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP608 | 38 | 140 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||



