 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | EMT12754 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Aegilops
|
||||||||
| Family | MIKC_MADS | ||||||||
| Protein Properties | Length: 312aa MW: 34689.9 Da PI: 10.0307 | ||||||||
| Description | MIKC_MADS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SRF-TF | 96.1 | 1.5e-30 | 9 | 59 | 1 | 51 |
S---SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-TTSEEEEEE- CS
SRF-TF 1 krienksnrqvtfskRrngilKKAeELSvLCdaevaviifsstgklyeyss 51
krien++ rqvtf+kRrng+lKKA+ELSvLCdaeva+i+fss+g+lyeys+
EMT12754 9 KRIENTTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYSN 59
79***********************************************95 PP
| |||||||
| 2 | K-box | 95.9 | 6.3e-32 | 144 | 235 | 9 | 100 |
K-box 9 leeakaeslqqelakLkkeienLqreqRhllGedLesLslkeLqqLeqqLekslkkiRskKnellleqieelqkkekelqeenkaLrkklee 100
e + ++++qqe+akL+ +i++Lq++++hl+G+++++LslkeL+qLe++Lek++ kiR++Knell ++i+++ k+e elq++ +Lr+k++e
EMT12754 144 IEVNAQQYYQQEAAKLRHQIQMLQSTNKHLVGDSVGNLSLKELKQLESRLEKGIAKIRARKNELLSSEINYMVKREIELQSDSIDLRTKIAE 235
5667899**********************************************************************************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50066 | 33.389 | 1 | 61 | IPR002100 | Transcription factor, MADS-box |
| SMART | SM00432 | 3.4E-41 | 1 | 60 | IPR002100 | Transcription factor, MADS-box |
| SuperFamily | SSF55455 | 9.42E-29 | 2 | 62 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00265 | 2.01E-38 | 2 | 60 | No hit | No description |
| PROSITE pattern | PS00350 | 0 | 3 | 57 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 9.4E-32 | 3 | 23 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF00319 | 2.3E-26 | 10 | 57 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 9.4E-32 | 23 | 38 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 9.4E-32 | 38 | 59 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF01486 | 1.9E-23 | 146 | 233 | IPR002487 | Transcription factor, K-box |
| PROSITE profile | PS51297 | 13.878 | 149 | 239 | IPR002487 | Transcription factor, K-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0048283 | Biological Process | indeterminate inflorescence morphogenesis | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0080155 | Biological Process | regulation of double fertilization forming a zygote and endosperm | ||||
| GO:0090376 | Biological Process | seed trichome differentiation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 312 aa Download sequence Send to blast |
MGRGRIEIKR IENTTSRQVT FCKRRNGLLK KAYELSVLCD AEVALIVFSS RGRLYEYSNN 60 RAVSQLGLGL AADCFCARRA SLTRGRARAS PPQPRPPLRV LQQQVTDVSL PVSPPLFINC 120 SVKATIDRYK KAHACGSTSG VPLIEVNAQQ YYQQEAAKLR HQIQMLQSTN KHLVGDSVGN 180 LSLKELKQLE SRLEKGIAKI RARKNELLSS EINYMVKREI ELQSDSIDLR TKIAEEEQRL 240 QQVTIARPSV APELNPFTAL DMKCFFPANL FEAAVHAHAQ AQAQAQAQAS LQLNLGYQQL 300 APPGAGDVAH QF |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5f28_A | 3e-17 | 1 | 59 | 1 | 59 | MEF2C |
| 5f28_B | 3e-17 | 1 | 59 | 1 | 59 | MEF2C |
| 5f28_C | 3e-17 | 1 | 59 | 1 | 59 | MEF2C |
| 5f28_D | 3e-17 | 1 | 59 | 1 | 59 | MEF2C |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
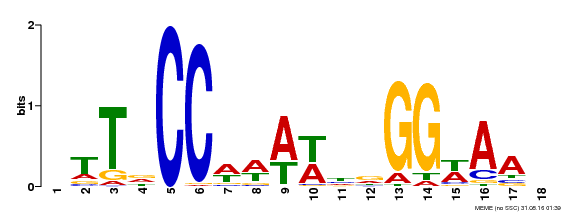
| MP00093 | SELEX | Transfer from AT3G58780 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | DQ512349 | 0.0 | DQ512349.1 Triticum aestivum MADS-box transcription factor TaAGL31 (AGL31) mRNA, complete cds. | |||
| GenBank | DQ512364 | 0.0 | DQ512364.1 Triticum aestivum MADS-box transcription factor TaAGL9 (AGL9) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020189538.1 | 1e-123 | MADS-box transcription factor 13-like | ||||
| Swissprot | Q2QW53 | 2e-85 | MAD13_ORYSJ; MADS-box transcription factor 13 | ||||
| TrEMBL | N1QX58 | 0.0 | N1QX58_AEGTA; MADS-box transcription factor 13 | ||||
| STRING | EMT12754 | 0.0 | (Aegilops tauschii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP649 | 37 | 144 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G58780.3 | 1e-72 | MIKC_MADS family protein | ||||



