 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | EMT12411 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Aegilops
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 358aa MW: 39424.4 Da PI: 8.2151 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 56.3 | 7.4e-18 | 92 | 139 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT+eEd +lv +++q+G+g+W+++++ g+ R+ k+c++rw +yl
EMT12411 92 KGPWTPEEDIILVSYIQQHGPGNWRSVPENTGLMRCSKSCRLRWTNYL 139
79********************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 46.5 | 8.4e-15 | 145 | 190 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg++T+ E+ +++++ ++lG++ W++Ia++++ Rt++++k++w+++l
EMT12411 145 RGNFTQHEEGIIIHLQALLGNK-WAAIASYLP-QRTDNDIKNYWNTHL 190
89********************.*********.************996 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 24.958 | 87 | 143 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 4.49E-30 | 89 | 186 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 7.4E-14 | 91 | 141 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.2E-16 | 92 | 139 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.7E-23 | 93 | 146 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.91E-12 | 94 | 139 | No hit | No description |
| SMART | SM00717 | 3.3E-13 | 144 | 192 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS51294 | 18.953 | 144 | 194 | IPR017930 | Myb domain |
| Pfam | PF00249 | 6.6E-13 | 145 | 190 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.4E-24 | 147 | 194 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.39E-9 | 147 | 190 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0001666 | Biological Process | response to hypoxia | ||||
| GO:0009617 | Biological Process | response to bacterium | ||||
| GO:0009626 | Biological Process | plant-type hypersensitive response | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0042761 | Biological Process | very long-chain fatty acid biosynthetic process | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 358 aa Download sequence Send to blast |
MDPRKEAAGC TLADFCMHGK HTSTGRRKAP YDMWSHAAEE SRRPASTAGL VSSNKRRPWG 60 EEKLSFRAIL SGGNDDMGRP PCCDNGGASV KKGPWTPEED IILVSYIQQH GPGNWRSVPE 120 NTGLMRCSKS CRLRWTNYLR PGIKRGNFTQ HEEGIIIHLQ ALLGNKWAAI ASYLPQRTDN 180 DIKNYWNTHL KKKVKRLQQQ THPADHFFQH TSTAPNAAEQ TTSMNYYNPS SSNLDSAMQA 240 AMGYPETTTG NVTPATMAVS KLFQSWMPKA PSTDLADYKG MVAMQEFRTD GEAGASVSMT 300 SGDRFSSSDI LTGRGKADMA TTFSVLENWL LDDMVPGQAA MDGLIDISAG CCANPIMF |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1gv2_A | 2e-25 | 92 | 194 | 4 | 105 | MYB PROTO-ONCOGENE PROTEIN |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in the regulation of gene (e.g. drought-regulated and flavonoid biosynthetic genes) expression and stomatal movements leading to negative regulation of responses to drought and responses to other physiological stimuli (e.g. light). {ECO:0000269|PubMed:22018045}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00386 | DAP | Transfer from AT3G28910 | Download |
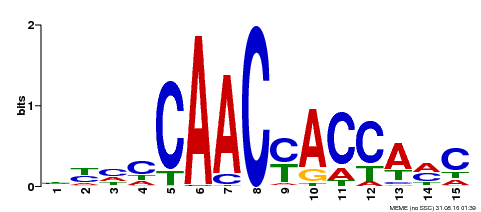
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Repressed by abscisic acid (ABA) and osmotic stress (salt stress). {ECO:0000269|PubMed:22018045}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK110844 | 1e-136 | AK110844.1 Oryza sativa Japonica Group cDNA clone:002-172-B09, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020177196.1 | 0.0 | transcription factor MYB30-like | ||||
| Swissprot | B3VTV7 | 1e-81 | MYB60_VITVI; Transcription factor MYB60 | ||||
| TrEMBL | M8BEL8 | 0.0 | M8BEL8_AEGTA; Myb-related protein 306 | ||||
| STRING | EMT12411 | 0.0 | (Aegilops tauschii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP229 | 38 | 296 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G28910.1 | 8e-79 | myb domain protein 30 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



