 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | EMT08990 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Aegilops
|
||||||||
| Family | AP2 | ||||||||
| Protein Properties | Length: 705aa MW: 73840.3 Da PI: 6.8916 | ||||||||
| Description | AP2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 34.8 | 4.1e-11 | 267 | 323 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrd.pseng..kr.krfslgkfgtaeeAakaaiaarkkleg 55
s y+GV++++++gr++A+++d + ++ k + g ++ +e+Aa+a++ a++k++g
EMT08990 267 SIYRGVTRHRWTGRYEAHLWDnSCR-RegQTrKGRQ-GGYDKEEKAARAYDLAALKYWG 323
57*******************4444.2455535555.779999**************98 PP
| |||||||
| 2 | AP2 | 46.7 | 8.1e-15 | 366 | 417 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s y+GV+++++ grW A+I + +k +lg+f t eeAa+a++ a+ k++g
EMT08990 366 SIYRGVTRHHQHGRWQARIGRVAG---NKDLYLGTFSTQEEAAEAYDIAAIKFRG 417
57****************988532...5************************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 1.5E-14 | 267 | 333 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 1.3E-8 | 267 | 323 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 2.26E-17 | 267 | 333 | No hit | No description |
| PROSITE profile | PS51032 | 17.28 | 268 | 331 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 1.4E-21 | 268 | 337 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 1.0E-11 | 268 | 332 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 4.7E-6 | 269 | 280 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 6.42E-23 | 366 | 425 | No hit | No description |
| Pfam | PF00847 | 1.0E-9 | 366 | 417 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 1.57E-17 | 366 | 426 | IPR016177 | DNA-binding domain |
| PROSITE profile | PS51032 | 19.045 | 367 | 425 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 2.4E-33 | 367 | 431 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 6.8E-18 | 367 | 425 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 4.7E-6 | 407 | 427 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000723 | Biological Process | telomere maintenance | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007389 | Biological Process | pattern specification process | ||||
| GO:0010449 | Biological Process | root meristem growth | ||||
| GO:0019827 | Biological Process | stem cell population maintenance | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 705 aa Download sequence Send to blast |
MATVNNWLGF SLSPQELPPS AAASGDVSGA DVCYNIPQDW NMRGSELSAL VAEPKLEDFL 60 GGISYSDHHH HQHKQPGGNN MVVPAGSASG GAACYASSGG SSVGYLYHPS SASLQFDDSG 120 MFADSVMVAS SGGGVHHDGA GIMANTTANG DLNNGGGGGG IGLSMIKSWL RSQPSPAQPP 180 EQQRADAAAQ GLSLSMNMAA CMPPLACGER GVPELAIVRK DDTAGGSGAG SGAVVSAGAD 240 STGGSSGAVV ETPASRKTAD TFGQRTSIYR GVTRHRWTGR YEAHLWDNSC RREGQTRKGR 300 QGGYDKEEKA ARAYDLAALK YWGATTTTNF PVNNYEKELE EMKHMTRQEF VASLRRKSSG 360 FSRGASIYRG VTRHHQHGRW QARIGRVAGN KDLYLGTFST QEEAAEAYDI AAIKFRGLNA 420 VTNFDMTRYD VKSILDSTAL PIGSAAKRLK DAEAATASSL ALQQQQHAGV VSGYDMGALA 480 AYGAAYHHHH PSVAGAVAWP TIAFQAPPQQ VSGGGGGHMY HPYAHAQPLR GWCKQEPDHA 540 VIAAAHSLQE LHHLNLGAGG GAHDFFSQHA QAMQQQHGGL GSIDNGAGAS LEHSTGSNSV 600 VYNNGAVESY MLPMSTTSTA AVSHAHEQAA AHAHARAVAA QAYSGGNMSR DHDDGKIAYE 660 NYLVSTAEAY GGGRMAAWAP ASAPPATSSS DMTGTQLFSV WNDTN |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00368 | DAP | Transfer from AT3G20840 | Download |
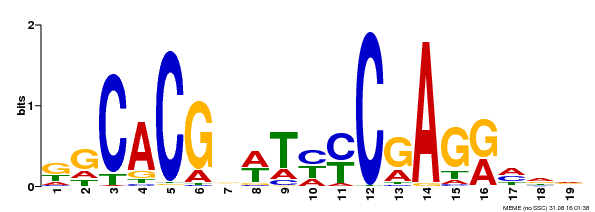
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HG670306 | 0.0 | HG670306.1 Triticum aestivum chromosome 3B, genomic scaffold, cultivar Chinese Spring. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020151353.1 | 0.0 | AP2-like ethylene-responsive transcription factor BBM2 | ||||
| TrEMBL | M8B5V2 | 0.0 | M8B5V2_AEGTA; AP2-like ethylene-responsive transcription factor BBM1 | ||||
| STRING | EMT08990 | 0.0 | (Aegilops tauschii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1569 | 37 | 110 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G17430.1 | 1e-117 | AP2 family protein | ||||



