 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | EMT03587 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Aegilops
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 403aa MW: 43803.9 Da PI: 10.1369 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 100.7 | 9.6e-32 | 261 | 316 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r++W+peLH+rFv+a++ LGG+++AtPk+i+elmkv+gLt+++vkSHLQkYRl+
EMT03587 261 KARRCWSPELHRRFVNALQILGGAQVATPKQIRELMKVDGLTNDEVKSHLQKYRLH 316
68****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 13.652 | 258 | 318 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.3E-28 | 259 | 319 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 2.24E-17 | 259 | 319 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.1E-26 | 261 | 316 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 8.0E-7 | 263 | 314 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009266 | Biological Process | response to temperature stimulus | ||||
| GO:0009740 | Biological Process | gibberellic acid mediated signaling pathway | ||||
| GO:0010452 | Biological Process | histone H3-K36 methylation | ||||
| GO:0048579 | Biological Process | negative regulation of long-day photoperiodism, flowering | ||||
| GO:1903507 | Biological Process | negative regulation of nucleic acid-templated transcription | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 403 aa Download sequence Send to blast |
MDSSPSDLTL DYTPNGNGTA SAGGGYPKQA SPVVDHHHHL SAEQATTQKL QEFLARLDDE 60 RLKIDAFKRE LPLCMQLLNQ AMEAYRQQLE ACQMGSHGGA AARAPLVLEE FIPLKNIGID 120 AAEKAAGNAP SEKANWMVSA QLWNGPAAGD AAAKGPQTPK ERSEHPLDTS PMLGALDGGG 180 GNGVGAFLPF TKEKACMAES AALPELGLAP AEKDREADRK PYHDDSGSSG SVVSRRDVGA 240 PPTSTAPEGQ SVPPPPQTNR KARRCWSPEL HRRFVNALQI LGGAQVATPK QIRELMKVDG 300 LTNDEVKSHL QKYRLHTRRP MPAPSAPPPG APPKKFVNSK MFMDVKKSSQ TSNKFTIGKR 360 IIDFKKKKFV NSKMFMDMKK SSQTSNKFTI GKRIIDFKKV HQI |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 8e-14 | 261 | 314 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 8e-14 | 261 | 314 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 7e-14 | 261 | 314 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 7e-14 | 261 | 314 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 7e-14 | 261 | 314 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 7e-14 | 261 | 314 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 8e-14 | 261 | 314 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 8e-14 | 261 | 314 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 8e-14 | 261 | 314 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 8e-14 | 261 | 314 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 8e-14 | 261 | 314 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 8e-14 | 261 | 314 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may play a role in regulatory networks controlling development and metabolism. {ECO:0000250|UniProtKB:Q6Z869}. | |||||
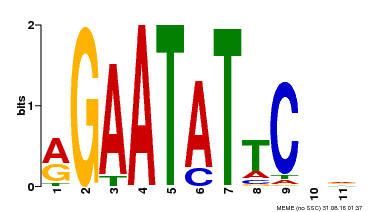
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00261 | DAP | Transfer from AT2G03500 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HG670306 | 0.0 | HG670306.1 Triticum aestivum chromosome 3B, genomic scaffold, cultivar Chinese Spring. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020184511.1 | 0.0 | myb family transcription factor EFM | ||||
| Swissprot | Q5VRW2 | 1e-114 | NOH1_ORYSJ; Transcription factor NIGTH1 | ||||
| TrEMBL | N1QTH5 | 0.0 | N1QTH5_AEGTA; Two-component response regulator ARR18 | ||||
| STRING | EMT03587 | 0.0 | (Aegilops tauschii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP9773 | 33 | 38 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G03500.1 | 2e-64 | G2-like family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



