 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 916752 | ||||||||
| Common Name | ARALYDRAFT_916752 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 611aa MW: 66771.6 Da PI: 4.9275 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 39.4 | 1.1e-12 | 433 | 478 | 4 | 54 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksL 54
+h e+Er+RR+++N++f Lr ++P+ K++Ka+ L A+ YI++L
916752 433 NHVEAERQRREKLNQRFYSLRAVVPNV-----SKMDKASLLGDAISYINEL 478
799***********************6.....5***************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 6.3E-55 | 56 | 239 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 17.282 | 429 | 478 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 3.01E-18 | 432 | 498 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 2.19E-14 | 432 | 483 | No hit | No description |
| Pfam | PF00010 | 4.0E-10 | 433 | 478 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 6.2E-18 | 433 | 497 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 9.8E-17 | 435 | 484 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006952 | Biological Process | defense response | ||||
| GO:0009718 | Biological Process | anthocyanin-containing compound biosynthetic process | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043425 | Molecular Function | bHLH transcription factor binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 611 aa Download sequence Send to blast |
MNGTTSSINF LTSDDDASAA MEAFIGTNHH SSLFPPPPQP PPPPSFPQTQ FNEDTLQQRL 60 QALIESAGEN WTYAIFWQIS HDFDSSTGDN TVILGWGDGY YKGEEDKEKK KNNTNTAEQE 120 HRKRVIRELN SLISGGIGVS DESNDEEVTD TEWFFLVSMT QSFVNGVGLP GESFLNSRVI 180 WLSGSGSLTG SGCERAGQGQ IYGLKTMVCI ATQNGVVELG SSEVISQSSD LMDKVNNLFN 240 FNNGGSGNNG GEASSWGFNL NPDQGENDPA LWISEPTNTG IESPARVNNG NNSNSNSKSD 300 SHQISKLDKN DISSVENQNR QSSCLVERDL NFSSSGLNQN GNFQGGLLKS NETLSFCGNE 360 SSKKRSPVSK GSNNDEGMLS FSTVVRSAAK SVDSDHSDLE ASVVKEAIVV EPPEKKPRKR 420 GRKPANGREE PLNHVEAERQ RREKLNQRFY SLRAVVPNVS KMDKASLLGD AISYINELKS 480 KLQQAESDKE EIQKKLDGMS KEGNNGKGGG SRAKERKSSN QDSTASSIEM EIDVKIIGWD 540 VMIRVQCSKK DHPGARFMEA LKELDLEVNH ASLSVVNDLM IQQATVKMGS QFFNHDQLKV 600 ALMTKVGENY * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4rru_A | 1e-139 | 5 | 235 | 1 | 227 | Transcription factor MYC3 |
| 4ywc_A | 1e-139 | 5 | 235 | 1 | 227 | Transcription factor MYC3 |
| 4ywc_B | 1e-139 | 5 | 235 | 1 | 227 | Transcription factor MYC3 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 414 | 422 | KKPRKRGRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in tryptophan, jasmonic acid (JA) and other stress-responsive gene regulation. With MYC2 and MYC4, controls additively subsets of JA-dependent responses. Can form complexes with all known glucosinolate-related MYBs to regulate glucosinolate biosynthesis. Binds to the G-box (5'-CACGTG-3') of promoters. Activates multiple TIFY/JAZ promoters. {ECO:0000269|PubMed:12136026, ECO:0000269|PubMed:21242320, ECO:0000269|PubMed:21321051, ECO:0000269|PubMed:21335373, ECO:0000269|PubMed:23943862}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00085 | PBM | Transfer from AT5G46760 | Download |
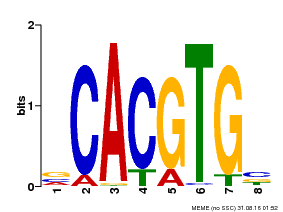
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 916752 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Barely up-regulated by jasmonic acid. {ECO:0000269|PubMed:21242320, ECO:0000269|PubMed:21335373}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB016882 | 0.0 | AB016882.1 Arabidopsis thaliana genomic DNA, chromosome 5, P1 clone:MZA15. | |||
| GenBank | CP002688 | 0.0 | CP002688.1 Arabidopsis thaliana chromosome 5 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002865164.1 | 0.0 | transcription factor MYC3 | ||||
| Swissprot | Q9FIP9 | 0.0 | MYC3_ARATH; Transcription factor MYC3 | ||||
| TrEMBL | D7MQP8 | 0.0 | D7MQP8_ARALL; Uncharacterized protein | ||||
| STRING | scaffold_800152.1 | 0.0 | (Arabidopsis lyrata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1265 | 28 | 96 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G46760.1 | 0.0 | bHLH family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 916752 |
| Entrez Gene | 9301238 |



