 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 886837 | ||||||||
| Common Name | ARALYDRAFT_686281 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | AP2 | ||||||||
| Protein Properties | Length: 443aa MW: 49282 Da PI: 7.1299 | ||||||||
| Description | AP2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 45 | 2.8e-14 | 176 | 235 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrd.pse.ng..kr.krfslgkfgtaeeAakaaiaarkkleg 55
s y+GV++++++gr++A+++d + g ++ ++++lg ++ ++ Aa+a++ a++k++g
886837 176 SVYRGVTRHRWTGRYEAHLWDnSCRrEGhaRKgRQVYLGGYDKEDRAARAYDLAALKYWG 235
78*******************666565598446**********99*************98 PP
| |||||||
| 2 | AP2 | 49.4 | 1.2e-15 | 280 | 329 | 3 | 55 |
AP2 3 ykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
y+GV++++++grW A+I + +k +lg+f t+eeAa+a++ a+ k++g
886837 280 YRGVTRHHQQGRWQARIGRVAG---NKDLYLGTFATEEEAAEAYDIAAIKFRG 329
9***************988532...5************************997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 1.05E-16 | 176 | 244 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 3.1E-11 | 176 | 235 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 5.12E-21 | 176 | 245 | No hit | No description |
| PROSITE profile | PS51032 | 19.085 | 177 | 243 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 1.7E-15 | 177 | 244 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 2.0E-26 | 177 | 249 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.3E-6 | 178 | 189 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 8.5E-18 | 278 | 338 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 8.95E-11 | 278 | 337 | No hit | No description |
| PROSITE profile | PS51032 | 19.269 | 279 | 337 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 3.3E-18 | 279 | 337 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 3.3E-30 | 279 | 343 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 5.6E-10 | 280 | 329 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.3E-6 | 319 | 339 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010492 | Biological Process | maintenance of shoot apical meristem identity | ||||
| GO:0060772 | Biological Process | leaf phyllotactic patterning | ||||
| GO:0060774 | Biological Process | auxin mediated signaling pathway involved in phyllotactic patterning | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 443 aa Download sequence Send to blast |
MADSTTLSTF FDSQIHSQAQ IPKLEDFLGD SFVRYSDNQT ETQDSSSLTR FYDLRHHTDA 60 GGVSEFFSDH HQQHDFKTIN SGSEIVDDST ASNIGGTHLS SHVVESSTTA ELGFNGGCTT 120 GGALSLGVNN TSDQHLISCN NGERGENSNK KKTVSKKETS DDSKKKTVET LGQRTSVYRG 180 VTRHRWTGRY EAHLWDNSCR REGHARKGRQ VYLGGYDKED RAARAYDLAA LKYWGPTATT 240 NFPVASYSKE LEEMNHMTKL EFIASLRRKS SGFSRGASMY RGVTRHHQQG RWQARIGRVA 300 GNKDLYLGTF ATEEEAAEAY DIAAIKFRGI NAVTNFEMNR YDVEAVMKSS FPVGGAAAKR 360 HKLSLESPSS SSSDHNIQQQ HLVPSSSSSD HNPNSIPCGI PFESSVLYHH QNFFQHYPLV 420 SDSTVQVPMN QAEFFLWPNQ SY* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probably acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. May be involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways (By similarity). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00589 | DAP | Transfer from AT5G65510 | Download |
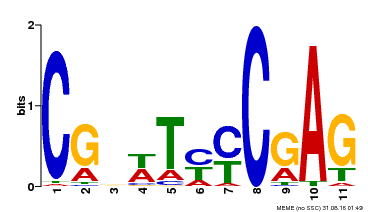
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 886837 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY560887 | 0.0 | AY560887.1 Arabidopsis thaliana putative AP2/EREBP transcription factor (At5g65510) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002864961.2 | 0.0 | AP2-like ethylene-responsive transcription factor AIL7 isoform X2 | ||||
| Swissprot | Q6J9N8 | 0.0 | AIL7_ARATH; AP2-like ethylene-responsive transcription factor AIL7 | ||||
| TrEMBL | D7MTZ1 | 0.0 | D7MTZ1_ARALL; Predicted protein | ||||
| STRING | Al_scaffold_0008_3101 | 0.0 | (Arabidopsis lyrata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3333 | 27 | 65 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G65510.1 | 0.0 | AINTEGUMENTA-like 7 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 886837 |
| Entrez Gene | 9302748 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



