 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 864075 | ||||||||
| Common Name | ARALYDRAFT_663519 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 343aa MW: 39529.1 Da PI: 6.9582 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 60.2 | 4.5e-19 | 15 | 62 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT+eEd++l+d+++++G g+W+t ++ g+ R++k+c++rw +yl
864075 15 KGPWTPEEDQKLIDYINKHGYGNWRTLPKNAGLQRCGKSCRLRWTNYL 62
79********************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 53.2 | 6.9e-17 | 68 | 112 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
rgr++ eE+e +++++ +G++ W++Ia++++ gRt++++k++w+++
864075 68 RGRFSFEEEETIIQLHSIMGNK-WSAIAARLP-GRTDNEIKNYWNTH 112
89********************.*********.************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.7E-25 | 7 | 65 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 25.462 | 10 | 66 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 3.34E-32 | 12 | 109 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 4.4E-14 | 14 | 64 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.5E-17 | 15 | 62 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.16E-12 | 17 | 62 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 1.1E-26 | 66 | 117 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.3E-16 | 67 | 115 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS51294 | 20.228 | 67 | 117 | IPR017930 | Myb domain |
| Pfam | PF00249 | 2.5E-15 | 68 | 112 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 7.23E-12 | 70 | 113 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 343 aa Download sequence Send to blast |
MGRSPCCEKK NGLKKGPWTP EEDQKLIDYI NKHGYGNWRT LPKNAGLQRC GKSCRLRWTN 60 YLRPDIKRGR FSFEEEETII QLHSIMGNKW SAIAARLPGR TDNEIKNYWN THIRKRLLKM 120 GIDPVTHTPR LDLLDISSIL SSSIYNYTHH HHHQHMNMSR LMMGDGHHQP LVNPEILKLA 180 TSLFSNQTHP NKTHENHTVN QTEVNQYQTG YNMTGNQELQ SWFPIMDQFA EFQDLMPMKT 240 TVQDSLPSFP SNHEDDYSKS NSVLEPYYSD FASVLTTPSS SPTPLNSSST TYINSSTCST 300 EDEKESYYSN NVMSTSEIAD YSFDVNGFLQ FQETKNNWDR VM* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 3e-29 | 13 | 117 | 5 | 108 | B-MYB |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may function in salt stress response. {ECO:0000305|PubMed:26139822}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00429 | DAP | Transfer from AT4G05100 | Download |
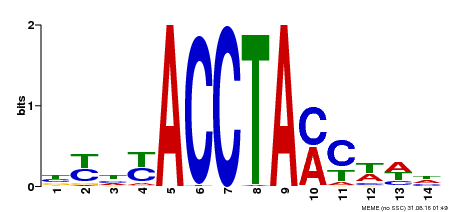
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 864075 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by salt stress. {ECO:0000269|PubMed:26139822}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF386932 | 0.0 | AF386932.1 Arabidopsis thaliana Unknown protein mRNA, complete cds. | |||
| GenBank | BT001182 | 0.0 | BT001182.1 Arabidopsis thaliana Unknown protein (At4g05100) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002872736.1 | 0.0 | transcription factor MYB74 | ||||
| Swissprot | Q9M0Y5 | 0.0 | MYB74_ARATH; Transcription factor MYB74 | ||||
| TrEMBL | D7M1Y3 | 0.0 | D7M1Y3_ARALL; Predicted protein | ||||
| STRING | Al_scaffold_0006_3584 | 0.0 | (Arabidopsis lyrata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4 | 28 | 2646 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G05100.1 | 1e-180 | myb domain protein 74 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 864075 |
| Entrez Gene | 9308805 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



