 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 487754 | ||||||||
| Common Name | ARALYDRAFT_487754 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 998aa MW: 112461 Da PI: 6.4036 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 187 | 1.9e-58 | 21 | 138 | 2 | 118 |
CG-1 2 lke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcywlLe 103
l+e ++rwl++ ei++iL+n++k+++++e++trp sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+hekLKvg+v+vl+cyYah+e n++fqrrcyw+Le
487754 21 LSEaQHRWLRPAEICEILQNYHKFHIASESPTRPASGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHEKLKVGSVDVLHCYYAHGEGNENFQRRCYWMLE 123
45559************************************************************************************************** PP
CG-1 104 eelekivlvhylevk 118
++l +iv+vhylevk
487754 124 QDLMHIVFVHYLEVK 138
************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 85.647 | 17 | 143 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 2.3E-85 | 20 | 138 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 2.9E-51 | 23 | 137 | IPR005559 | CG-1 DNA-binding domain |
| SuperFamily | SSF81296 | 1.23E-14 | 402 | 488 | IPR014756 | Immunoglobulin E-set |
| Pfam | PF01833 | 2.6E-4 | 402 | 477 | IPR002909 | IPT domain |
| Gene3D | G3DSA:2.60.40.10 | 2.7E-4 | 403 | 477 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF12796 | 1.5E-7 | 584 | 662 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 19.837 | 585 | 696 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 3.4E-18 | 585 | 697 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 3.58E-19 | 586 | 696 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.48E-14 | 591 | 694 | No hit | No description |
| SMART | SM00248 | 2900 | 602 | 631 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 8.63 | 602 | 634 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 11.22 | 635 | 667 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.0029 | 635 | 664 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 8.23E-8 | 810 | 860 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.51 | 810 | 832 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 7.73 | 811 | 840 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0024 | 813 | 831 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0092 | 833 | 855 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.395 | 834 | 858 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 4.4E-4 | 836 | 855 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0050826 | Biological Process | response to freezing | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 998 aa Download sequence Send to blast |
MVDRGSFGFI SPPQLDMEQL LSEAQHRWLR PAEICEILQN YHKFHIASES PTRPASGSLF 60 LFDRKVLRYF RKDGHNWRKK KDGKTVKEAH EKLKVGSVDV LHCYYAHGEG NENFQRRCYW 120 MLEQDLMHIV FVHYLEVKGN RTSIGMKENN SNSVNGTASV NIDSTASPTS TLSSLCEDAD 180 TGDSHQASSV LRASSEPQTG NRYGWTPAPG MRNVSQVHGN RVRESDSQRL VDVRAWDAIG 240 NSVTRYHDQP YCNNLLTQMQ PSNTDSMLVE ENTDKGGRLK AEHIRNPLQT QLNWQQNAQY 300 NFETFSSLLG SENQQPFGIS YQAPPSSMES EFIPVKKSLL RSEESLKKVD SFSRWASKEL 360 GEMEDLQMQS SRGDIAWTTV ECETAAAGIS LSPSLSEDQR FTIVDFWPKC AQTDAEVEVM 420 VIGTFLLSPQ EVTKYNWSCM FGEVEVPAEI LVDGVLCCHA PPHTAGHVPF YVTCSNRFAC 480 SEVREFDFLS GSTQKIDATD VYGTYTNEAS LQLRFEKMLA HRNFVHEHHI FKGVGEKRRK 540 ISKIMSLKEE KEYLLPGTYQ RDSTKQEPKE QLFREQSEEE LYIWLIHKVT EEGKGPNILD 600 EDGQGILHFV AALGYDWAIK PMLAAGVNIN FRDANGWSAL HWAAFSGREE TVAVLVSLGA 660 DAGALTDPSP ELPLGKTAAD LAYANGHRGI SGFLAESSLT SYLEKLTVDS KENSPANSSG 720 AKAVQTVSER TAAPMSYGDV PEKLSLKDSL TAVRNATQAA DRLHQVFRMQ SFQRKQLSDI 780 GDDDKIDISD KLAVSFATLK TKNLGQGDVS LSSAATHIQK KYRGWKKRKE FLLIRQRIVK 840 IQAHVRGHQV RKQYRTVIWS VGLLEKIILR WRRKGNGLRG FKRNAVAKTV EPEPPVSAIC 900 PTIPQEDEYD YLKEGRKQTE ERLEKALTRV KSMVQYPEAR DQYRRLLTVV EGFRENEASS 960 SASINNKEED EVNCEEDEFI DIDSLLNDDT LMMSISP* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds calmodulin in a calcium-dependent manner in vitro (PubMed:11925432). Binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Involved in freezing tolerance (PubMed:19270186). Involved in freezing tolerance in association with CAMTA2 and CAMTA3. Contributes together with CAMTA2 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved in drought stress responses by regulating several drought-responsive genes (PubMed:23547968). Involved in auxin signaling and responses to abiotic stresses (PubMed:20383645). Activates the expression of the V-PPase proton pump AVP1 in pollen (PubMed:14581622). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:11925432, ECO:0000269|PubMed:14581622, ECO:0000269|PubMed:19270186, ECO:0000269|PubMed:20383645, ECO:0000269|PubMed:23547968, ECO:0000269|PubMed:23581962, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00501 | DAP | Transfer from AT5G09410 | Download |
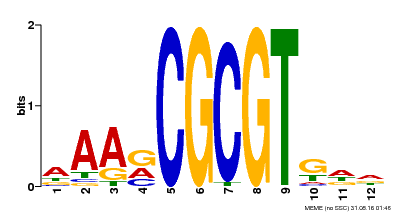
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 487754 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by UVB, wounding, ethylene and methyl jasmonate (PubMed:12218065). Induced by salt stress and heat shock (PubMed:12218065, PubMed:20383645). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:20383645, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK228740 | 0.0 | AK228740.1 Arabidopsis thaliana mRNA for Calmodulin-binding transcription activator 1, complete cds, clone: RAFL16-07-H14. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020878491.1 | 0.0 | calmodulin-binding transcription activator 1 isoform X4 | ||||
| Swissprot | Q9FY74 | 0.0 | CMTA1_ARATH; Calmodulin-binding transcription activator 1 | ||||
| TrEMBL | D7M234 | 0.0 | D7M234_ARALL; Calmodulin-binding transcription activator 1 | ||||
| STRING | fgenesh2_kg.6__888__AT5G09410.2 | 0.0 | (Arabidopsis lyrata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4082 | 24 | 52 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G09410.2 | 0.0 | ethylene induced calmodulin binding protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 487754 |
| Entrez Gene | 9307439 |



