 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 487412 | ||||||||
| Common Name | ARALYDRAFT_487412 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 520aa MW: 58094 Da PI: 6.091 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 54 | 3.9e-17 | 34 | 81 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+W+ Ed++l+d+v+++G g+W+++ r+ g+ R++k+c++rw ++l
487412 34 KGPWSSAEDDILIDYVNKHGEGNWNAVQRHTGLFRCGKSCRLRWANHL 81
79******************************************9986 PP
| |||||||
| 2 | Myb_DNA-binding | 52.3 | 1.3e-16 | 87 | 130 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
+g++++eE++l++++++++G++ W++ a++++ gRt++++k++w++
487412 87 KGAFSQEEEQLILELHAKMGNR-WARMAAHLP-GRTDNEIKNYWNT 130
799*******************.*********.***********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF001693 | 3.4E-280 | 1 | 519 | IPR016310 | Transcription factor, GAMYB |
| PROSITE profile | PS51294 | 15.406 | 29 | 81 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 7.11E-30 | 32 | 128 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 8.4E-15 | 33 | 83 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.2E-15 | 34 | 81 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 4.2E-23 | 35 | 88 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.91E-11 | 36 | 81 | No hit | No description |
| PROSITE profile | PS51294 | 26.222 | 82 | 136 | IPR017930 | Myb domain |
| SMART | SM00717 | 2.4E-17 | 86 | 134 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.2E-15 | 87 | 130 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.15E-12 | 89 | 130 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 1.1E-25 | 89 | 135 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009789 | Biological Process | positive regulation of abscisic acid-activated signaling pathway | ||||
| GO:0043068 | Biological Process | positive regulation of programmed cell death | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0045926 | Biological Process | negative regulation of growth | ||||
| GO:0048235 | Biological Process | pollen sperm cell differentiation | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 520 aa Download sequence Send to blast |
MSYTSTDSDH NESPVADDNG SDCRSRWEGH ALKKGPWSSA EDDILIDYVN KHGEGNWNAV 60 QRHTGLFRCG KSCRLRWANH LRPNLKKGAF SQEEEQLILE LHAKMGNRWA RMAAHLPGRT 120 DNEIKNYWNT RIKRRQRAGL PLYPPEMHVE ALDWSQEYAK SRVMGEDGRH QDFLQLGSCE 180 SNFFFDSLNF TDMVPGAFDL ADMTAYKNLG NGASSPRYEN FMTPIMPSSK RLWESELLYP 240 GCSSTVKQEF LSPEQFQNTS PQKISKTCSF SVPCDVEHPL YGNRHSPIMI SDSHTPTDGI 300 VPSSKPSYGA VKLELPSFQY SETTFDQWKK SSSPPHSHLL DPFDTYIQSP PPPTGREESD 360 LYSSFDTGLL DMLLLEAKIR NNTTKNNLYK SCASTIPSAD LGKVTVSQTK SEDFDNSLKS 420 LVHSDMSTQN ADGIPPRQRE KKRKPLLDIT RPDVLLASSW LDHGLGIVKE TGSMSDALTV 480 LLGDDIGNDY MNMSVGASSG VGSCSWSNMP PVCQMTELP* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1h8a_C | 7e-31 | 12 | 135 | 2 | 127 | MYB TRANSFORMING PROTEIN |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 435 | 443 | PRQREKKRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator of alpha-amylase expression that binds to 5'-CAACTGTC-3' motif in target gene promoter (PubMed:11743113). Positive regulator of abscisic acid (ABA) responses leading to growth arrest during seed germination (PubMed:17217461). In vegetative tissues, inhibits growth by reducing cell proliferation. Promotes the expression of aleurone-related genes (e.g. CP1, CP, GASA1, BXL1 and BXL2) in seeds. Together with MYB65 and MYB101, promotes the programmed cell death (PCD) the vacuolation of protein storage vacuoles (PSVs) in the aleurone layers during seed germination (PubMed:20699403). Binds to a GARE site (GA-response element) in the LEAFY promoter, essential for its gibberellic acid (GA)-mediated induction (PubMed:15226253). Together with MYB65, facilitates anther and tapetum development (PubMed:15722475). {ECO:0000269|PubMed:11743113, ECO:0000269|PubMed:15226253, ECO:0000269|PubMed:15722475, ECO:0000269|PubMed:17217461, ECO:0000269|PubMed:20699403}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00490 | DAP | Transfer from AT5G06100 | Download |
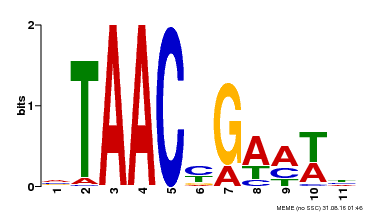
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 487412 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Accumulates at the shoot apex upon the transition from short- to long-day photoperiods leading to flowering and after gibberellins (GAs) treatment (PubMed:11743113). Repressed by microRNA159 (miR159a and miR159b) in vegetative tissues (PubMed:15226253, PubMed:20699403, PubMed:17916625). Specific expression in floral organs and in the shoot apices is regulated via miR159-mediated degradation (PubMed:15722475). Repressed in germinating seeds by miR159-mediated cleavage in an abscisic acid (ABA) and ABI3-dependent manner, probably to desensitize hormone signaling during seedling stress responses (PubMed:17217461, PubMed:18305205). Slightly induced by ethylene and cytokinins (PubMed:9839469). {ECO:0000269|PubMed:11743113, ECO:0000269|PubMed:15226253, ECO:0000269|PubMed:15722475, ECO:0000269|PubMed:17217461, ECO:0000269|PubMed:17916625, ECO:0000269|PubMed:18305205, ECO:0000269|PubMed:20699403, ECO:0000269|PubMed:9839469}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF411969 | 0.0 | AF411969.1 Arabidopsis thaliana putative transcription factor MYB33 (MYB33) mRNA, complete cds. | |||
| GenBank | AK118937 | 0.0 | AK118937.1 Arabidopsis thaliana At5g06100 mRNA for putative transcription factor MYB33, complete cds, clone: RAFL21-27-I01. | |||
| GenBank | AY519616 | 0.0 | AY519616.1 Arabidopsis thaliana MYB transcription factor (At5g06100) mRNA, complete cds. | |||
| GenBank | BT006049 | 0.0 | BT006049.1 Arabidopsis thaliana clone U51311 putative MYB family transcription factor (At5g06100) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020875964.1 | 0.0 | transcription factor MYB33 | ||||
| Refseq | XP_020875965.1 | 0.0 | transcription factor MYB33 | ||||
| Swissprot | Q8W1W6 | 0.0 | MYB33_ARATH; Transcription factor MYB33 | ||||
| TrEMBL | D7LZ38 | 0.0 | D7LZ38_ARALL; ATMYB33/MYB33 | ||||
| STRING | fgenesh2_kg.6__546__AT5G06100.3 | 0.0 | (Arabidopsis lyrata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3802 | 28 | 59 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06100.3 | 0.0 | myb domain protein 33 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 487412 |
| Entrez Gene | 9307268 |



