 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 477989 | ||||||||
| Common Name | ARALYDRAFT_477989 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 138aa MW: 15217.3 Da PI: 10.4797 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 56.8 | 3e-18 | 31 | 65 | 1 | 35 |
GATA 1 CsnCgttkTplWRrgpdgnktLCnaCGlyyrkkgl 35
C+ Cgt+kTplWR gp g+k+LCnaCG+++rkk++
477989 31 CAICGTSKTPLWRGGPAGPKSLCNACGIRNRKKRR 65
999*****************************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50114 | 12.387 | 25 | 61 | IPR000679 | Zinc finger, GATA-type |
| SMART | SM00401 | 9.9E-14 | 25 | 77 | IPR000679 | Zinc finger, GATA-type |
| SuperFamily | SSF57716 | 1.19E-12 | 27 | 73 | No hit | No description |
| Gene3D | G3DSA:3.30.50.10 | 8.4E-15 | 29 | 65 | IPR013088 | Zinc finger, NHR/GATA-type |
| CDD | cd00202 | 2.21E-12 | 30 | 85 | No hit | No description |
| PROSITE pattern | PS00344 | 0 | 31 | 56 | IPR000679 | Zinc finger, GATA-type |
| Pfam | PF00320 | 2.7E-16 | 31 | 65 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 138 aa Download sequence Send to blast |
MESKLTSVDA IEEHSSSSSS NEGISNEKKS CAICGTSKTP LWRGGPAGPK SLCNACGIRN 60 RKKRRTLISN RSEDKKNKNH NRNPKFGDSL KQRLMELGRE VMMQRSTAEN QRRKKLGEEE 120 QAAVLLMALS YASSVYA* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional regulator that specifically binds 5'-GATA-3' or 5'-GAT-3' motifs within gene promoters. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00335 | DAP | Transfer from AT3G06740 | Download |
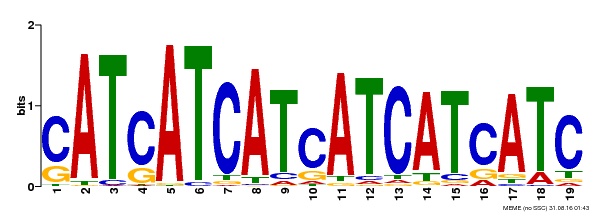
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 477989 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY063926 | 0.0 | AY063926.1 Arabidopsis thaliana unknown protein (At3g06740) mRNA, complete cds. | |||
| GenBank | AY091250 | 0.0 | AY091250.1 Arabidopsis thaliana unknown protein (At3g06740) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020888043.1 | 5e-97 | GATA transcription factor 15 | ||||
| Swissprot | Q8LG10 | 6e-89 | GAT15_ARATH; GATA transcription factor 15 | ||||
| TrEMBL | D7L5Z2 | 2e-95 | D7L5Z2_ARALL; Uncharacterized protein | ||||
| STRING | fgenesh2_kg.3__690__AT3G06740.1 | 4e-96 | (Arabidopsis lyrata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM5050 | 26 | 52 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G06740.1 | 1e-78 | GATA transcription factor 15 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 477989 |
| Entrez Gene | 9318557 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



