 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 476042 | ||||||||
| Common Name | ARALYDRAFT_476042 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | YABBY | ||||||||
| Protein Properties | Length: 182aa MW: 19750.5 Da PI: 10.0212 | ||||||||
| Description | YABBY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | YABBY | 137.6 | 1.4e-42 | 16 | 156 | 5 | 163 |
YABBY 5 ssseqvCyvqCnfCntilavsvPstslfkvvtvrCGhCtsllsvnlakasqllaaeshldeslkeelleelkveeenlksnvekeesastsvsseklsenede 107
++e++ yv+C+ Cntilav +P + ++ +vtv+CGhC +l ++ + + + h + l + +++ + k+ s+s+s ss +++++
476042 16 PQTEHLYYVRCSICNTILAVGIPLKRMLDTVTVKCGHCGNLSFLTT-----TPPLQGH--------VS--LTLQMQSFGGSEYKKGSSSSSSSS---TSSDQP 100
679************************************9755332.....2333333........32..333445566666666666666555...345666 PP
YABBY 108 evprvppvirPPekrqrvPsaynrfikeeiqrikasnPdishreafsaaaknWahf 163
+p +p v++PPek+qr Psaynrf+++eiqrik++nP+i hreafsaaaknWa +
476042 101 PSPTPPFVVKPPEKKQRLPSAYNRFMRDEIQRIKSANPEIPHREAFSAAAKNWAKY 156
7787788***********************************************86 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF04690 | 1.1E-49 | 18 | 156 | IPR006780 | YABBY protein |
| SuperFamily | SSF47095 | 1.27E-8 | 100 | 155 | IPR009071 | High mobility group box domain |
| Gene3D | G3DSA:1.10.30.10 | 7.1E-5 | 110 | 155 | IPR009071 | High mobility group box domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010254 | Biological Process | nectary development | ||||
| GO:0010582 | Biological Process | floral meristem determinacy | ||||
| GO:0048479 | Biological Process | style development | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 182 aa Download sequence Send to blast |
MNLEEKPTMA ASRASPQTEH LYYVRCSICN TILAVGIPLK RMLDTVTVKC GHCGNLSFLT 60 TTPPLQGHVS LTLQMQSFGG SEYKKGSSSS SSSSTSSDQP PSPTPPFVVK PPEKKQRLPS 120 AYNRFMRDEI QRIKSANPEI PHREAFSAAA KNWAKYIPNS PTSITSGGHN MIHGLGFGEK 180 K* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor required for the initiation of nectary development. Also involved in suppressing early radial growth of the gynoecium, in promoting its later elongation and in fusion of its carpels by regulating both cell division and expansion. Establishes the polar differentiation in the carpels by specifying abaxial cell fate in the ovary wall. Regulates both cell division and expansion. {ECO:0000269|PubMed:10225997, ECO:0000269|PubMed:10225998, ECO:0000269|PubMed:10535738, ECO:0000269|PubMed:11714690, ECO:0000269|PubMed:15598802, ECO:0000269|Ref.10}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
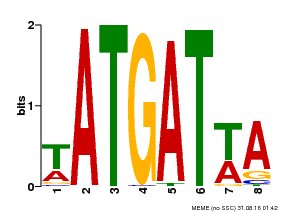
| Motif ID | Method | Source | Motif file |
| MP00220 | DAP | Transfer from AT1G69180 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 476042 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated by SPT and by A class genes AP2 and LUG in the outer whorl. In the third whorl, B class genes AP3 and PI, and the C class gene AG act redundantly with each other and in combination with SEP1, SEP2, SEP3, SHP1 and SHP2 to activate CRC in nectaries and carpels. LFY enhances its expression. {ECO:0000269|PubMed:10225998, ECO:0000269|PubMed:15598802}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF132606 | 0.0 | AF132606.1 Arabidopsis thaliana transcription factor CRC mRNA, complete cds. | |||
| GenBank | AK229609 | 0.0 | AK229609.1 Arabidopsis thaliana mRNA for transcription factor CRC, complete cds, clone: RAFL19-52-F14. | |||
| GenBank | BT008618 | 0.0 | BT008618.1 Arabidopsis thaliana clone RAFL19-52-F14 (R50809) putative transcription factor CRC (At1g69180) mRNA, complete cds. | |||
| GenBank | DQ446412 | 0.0 | DQ446412.1 Arabidopsis thaliana clone pENTR221-At1g69180 transcription factor CRC (At1g69180) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002888705.1 | 1e-133 | protein CRABS CLAW | ||||
| Swissprot | Q8L925 | 1e-116 | CRC_ARATH; Protein CRABS CLAW | ||||
| TrEMBL | D7KWZ0 | 1e-132 | D7KWZ0_ARALL; Uncharacterized protein | ||||
| STRING | fgenesh2_kg.2__1172__AT1G69180.1 | 1e-133 | (Arabidopsis lyrata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM9725 | 28 | 35 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G69180.1 | 1e-117 | YABBY family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 476042 |
| Entrez Gene | 9323290 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



