 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 471409 | ||||||||
| Common Name | ARALYDRAFT_471409 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 200aa MW: 22368 Da PI: 5.1459 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 47.6 | 4.2e-15 | 18 | 68 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkr..krfslgkfgtaeeAakaaiaarkkleg 55
+ y+G+r++ +Wv+e+r+p + +r++lg++ ta++Aa+a++ a ++l+g
471409 18 PVYRGIRRRN-GDKWVCEVREP-----ThqRRIWLGTYPTADMAARAHDVAVLALRG 68
68*****877.9*********8.....336*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF00847 | 2.4E-10 | 18 | 68 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 3.33E-19 | 19 | 78 | IPR016177 | DNA-binding domain |
| Gene3D | G3DSA:3.30.730.10 | 2.3E-28 | 19 | 78 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 2.7E-23 | 19 | 82 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 21.562 | 19 | 76 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 2.86E-25 | 20 | 78 | No hit | No description |
| PRINTS | PR00367 | 3.0E-7 | 20 | 31 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.0E-7 | 42 | 58 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009408 | Biological Process | response to heat | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0010371 | Biological Process | regulation of gibberellin biosynthetic process | ||||
| GO:0016049 | Biological Process | cell growth | ||||
| GO:0048510 | Biological Process | regulation of timing of transition from vegetative to reproductive phase | ||||
| GO:0050826 | Biological Process | response to freezing | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 200 aa Download sequence Send to blast |
MRPKKRAGRR VFKETRHPVY RGIRRRNGDK WVCEVREPTH QRRIWLGTYP TADMAARAHD 60 VAVLALRGRS ACLNFADSAW QLPVPESNDP DVIRRVAAEA AEMFRPVDLG SGITVLPSLG 120 DDVDLGFGSG SGSGSGSEER NSSSYAFGDY EEVSTTMMRL AEEPLMSPPR SYTEDMTPSN 180 VYTEEEMCYE DISLWSYSY* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-[AG]CCGAC-3'. Binding to the C-repeat/DRE element mediates cold or dehydration-inducible transcription. CBF/DREB1 factors play a key role in freezing tolerance and cold acclimation. {ECO:0000269|PubMed:11798174}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00001 | PBM | 25378322 | Download |
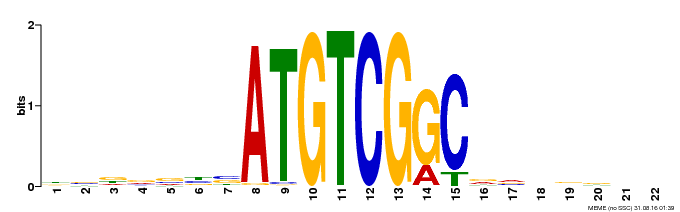
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 471409 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By high-salt stress. {ECO:0000269|PubMed:11798174}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC025417 | 0.0 | AC025417.4 Genomic sequence for Arabidopsis thaliana BAC T12C24 from chromosome I, complete sequence. | |||
| GenBank | AK228194 | 0.0 | AK228194.1 Arabidopsis thaliana mRNA for hypothetical protein, complete cds, clone: RAFL14-63-M15. | |||
| GenBank | AY560892 | 0.0 | AY560892.1 Arabidopsis thaliana putative AP2/EREBP transcription factor (At1g12610) mRNA, complete cds. | |||
| GenBank | CP002684 | 0.0 | CP002684.1 Arabidopsis thaliana chromosome 1 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020870917.1 | 1e-146 | dehydration-responsive element-binding protein 1F | ||||
| Swissprot | Q9LN86 | 1e-131 | DRE1F_ARATH; Dehydration-responsive element-binding protein 1F | ||||
| TrEMBL | D7KP85 | 1e-145 | D7KP85_ARALL; Uncharacterized protein | ||||
| STRING | fgenesh2_kg.1__1363__AT1G12610.1 | 1e-146 | (Arabidopsis lyrata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM355 | 28 | 187 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G12610.1 | 1e-124 | ERF family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 471409 |
| Entrez Gene | 9328759 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



