 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 327841 | ||||||||
| Common Name | ARALYDRAFT_327841 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | BBR-BPC | ||||||||
| Protein Properties | Length: 445aa MW: 49570.8 Da PI: 9.1087 | ||||||||
| Description | BBR-BPC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GAGA_bind | 230.8 | 1.1e-70 | 22 | 279 | 8 | 272 |
GAGA_bind 8 e..rnkgyye................paaslkenlglqlmssiaerdakirernlalsekkaavaerdmaflqrdkalaernkalverdnkllalllven 89
e r k+ y+ ++++l +++++ms++aerda+++e n a++++k+a+a+rd+a+ qrdkal+er++a++e +++l al ++en
327841 22 EngRYKPDYNkgtqsvnmmpqhqikeQHNALV--MNKKIMSILAERDAAVKEINEAVAATKEALAARDEALEQRDKALSERDNAIMETESALNALRYREN 119
44566664444666666666555543556666..99**************************************************************** PP
GAGA_bind 90 slasalpvgvqvlsgtksidslqqlsepqledsavelreeeklealpieeaaeeakekkkkkkrqrakkpkekkakkkkkksekskkkvkkesaderska 189
+l+ l+ + +g ++ + e++ pi++ ++ea ++k k r+k+ k+ ++k kk +e+ +++v++++ ++s++
327841 120 NLNYILS--CA-KRGGSHSC---------------VTDESHLPAPSPISTIPPEAAHTKLAK---RKKESKQGARSKGKKVGEDLNRQVASPG--KKSRK 196
9664441..11.12222221...............123455567789999999998888887...55666666677777888899********..56*** PP
GAGA_bind 190 ekksidlvlngvslDestlPvPvCsCtGalrqCYkWGnGGWqSaCCtttiSvyPLPvstkrrgaRiagrKmSqgafkklLekL 272
+++s d++ln v++De+t+PvP+C+CtG+ rqCYkWGnGGWqS+CCttt+S+yPLP+++++r++R++grKmS+++f++lL+ L
327841 197 DWDSYDVGLNLVTFDETTMPVPMCTCTGTARQCYKWGNGGWQSSCCTTTLSQYPLPQMPNKRHSRVGGRKMSGSVFSRLLSLL 279
********************************************************************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01226 | 2.0E-137 | 15 | 282 | IPR010409 | GAGA-binding transcriptional activator |
| Pfam | PF06217 | 1.3E-67 | 45 | 279 | IPR010409 | GAGA-binding transcriptional activator |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016021 | Cellular Component | integral component of membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 445 aa Download sequence Send to blast |
MFDIILAIAV ESKRMECGGQ YENGRYKPDY NKGTQSVNMM PQHQIKEQHN ALVMNKKIMS 60 ILAERDAAVK EINEAVAATK EALAARDEAL EQRDKALSER DNAIMETESA LNALRYRENN 120 LNYILSCAKR GGSHSCVTDE SHLPAPSPIS TIPPEAAHTK LAKRKKESKQ GARSKGKKVG 180 EDLNRQVASP GKKSRKDWDS YDVGLNLVTF DETTMPVPMC TCTGTARQCY KWGNGGWQSS 240 CCTTTLSQYP LPQMPNKRHS RVGGRKMSGS VFSRLLSLLT IRFKSRMKDE ARFVIVFTVD 300 SSVNANILVL VFFATFFSGI AFAFEWTFHG KNHSGFQWII YYALSLIILP ILIWLGLGIV 360 MIVSSSHGSM QVASVAVEEE QCVNSFAEDS GNEETKENNC ESLAIVVDSD KKKCTDKVFE 420 NKTAKLKRTV SFPLHSHVRS CRTR* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional regulator that specifically binds to GA-rich elements (GAGA-repeats) present in regulatory sequences of genes involved in developmental processes. {ECO:0000269|PubMed:14731261}. | |||||
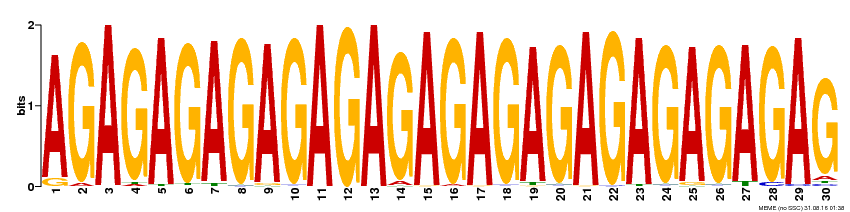
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00480 | DAP | Transfer from AT4G38910 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 327841 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | EF633603 | 0.0 | EF633603.1 Cardamine pratensis GAGA-motif binding transcriptional activator (BBR/BPC5) gene, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020875421.1 | 0.0 | protein BASIC PENTACYSTEINE5 isoform X1 | ||||
| Swissprot | F4JUI3 | 1e-165 | BPC5_ARATH; Protein BASIC PENTACYSTEINE5 | ||||
| TrEMBL | D7MFP8 | 0.0 | D7MFP8_ARALL; Uncharacterized protein | ||||
| STRING | fgenesh1_pm.C_scaffold_7000068 | 0.0 | (Arabidopsis lyrata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4090 | 26 | 59 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G38910.2 | 1e-160 | basic pentacysteine 5 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 327841 |
| Entrez Gene | 9304943 |



