 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Araip.Y4IW0 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Dalbergieae; Arachis
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 100aa MW: 11414.6 Da PI: 9.471 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 139.8 | 7.6e-44 | 19 | 95 | 2 | 78 |
-SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 2 CqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
Cqv+gC++dlseak+yhrrhkvCe+h+kap+v +sg++qrfCqqCsrfhelsefD++krsCrrrLa+hnerrrk+++
Araip.Y4IW0 19 CQVDGCDSDLSEAKQYHRRHKVCEFHAKAPSVHISGQQQRFCQQCSRFHELSEFDDSKRSCRRRLAGHNERRRKNAT 95
**************************************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 1.8E-37 | 11 | 80 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 33.112 | 16 | 93 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 3.27E-40 | 17 | 97 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 3.9E-35 | 19 | 92 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0010229 | Biological Process | inflorescence development | ||||
| GO:0010321 | Biological Process | regulation of vegetative phase change | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 100 aa Download sequence Send to blast |
MSSKRGFKGG GGSSMAPSCQ VDGCDSDLSE AKQYHRRHKV CEFHAKAPSV HISGQQQRFC 60 QQCSRFHELS EFDDSKRSCR RRLAGHNERR RKNATEYHGE |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 8e-39 | 9 | 92 | 1 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Binds specifically to the 5'-GTAC-3' core sequence. Promotes both vegetative phase change and flowering. Regulates phase-specific patterns of leaf epidermal differentiation and flowering time, but does not seem to affect leaf shape. {ECO:0000269|PubMed:16095614, ECO:0000269|PubMed:16914499, ECO:0000269|PubMed:9301089}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00290 | DAP | Transfer from AT2G33810 | Download |
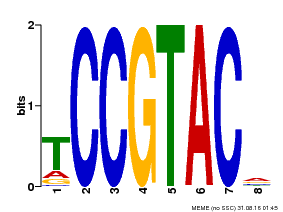
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Araip.Y4IW0 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Negatively regulated by microRNAs miR156. {ECO:0000269|PubMed:12202040, ECO:0000269|PubMed:16914499}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KF312225 | 1e-155 | KF312225.1 Arachis hypogaea squamosa promoter-binding-like protein (SPL3) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_016190070.1 | 6e-69 | squamosa promoter-binding-like protein 3 isoform X2 | ||||
| Refseq | XP_025638265.1 | 6e-69 | squamosa promoter-binding-like protein 3 isoform X1 | ||||
| Swissprot | P93015 | 4e-41 | SPL3_ARATH; Squamosa promoter-binding-like protein 3 | ||||
| TrEMBL | A0A444ZWG3 | 2e-67 | A0A444ZWG3_ARAHY; Squamosa promoter-binding-like protein | ||||
| STRING | GLYMA13G31090.1 | 8e-52 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF535 | 34 | 153 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G33810.1 | 4e-30 | squamosa promoter binding protein-like 3 | ||||



