 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Araip.PF1MQ | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Dalbergieae; Arachis
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 861aa MW: 93932.9 Da PI: 6.4458 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 66.1 | 4.9e-21 | 157 | 212 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+k+l L++rqVk+WFqNrR+++k
Araip.PF1MQ 157 KKRYHRHTPQQIQELEALFKECPHPDEKQRLELSKRLCLETRQVKFWFQNRRTQMK 212
688999***********************************************999 PP
| |||||||
| 2 | START | 202.4 | 1.9e-63 | 376 | 602 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE.....EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEEEEE CS
START 1 elaeeaaqelvkkalaeepgWvkss.....esengdevlqkfeeskv.....dsgealrasgvvdmvlallveellddkeqWdetla....kaetlev 84
ela+ a++elvk+a+++ep+W + e +n +e+ +++++ + + +ea+r++++v+ ++ lv +l+d++ +W+e+++ + +t+ev
Araip.PF1MQ 376 ELALGAMDELVKMAQSGEPLWIRGMemggrEVLNHEEYSRTVTPCIGlrpngFVSEASRETAIVIINSLPLVDTLMDSN-RWSEMFPcmvaRTSTTEV 472
57899*****************99989999999999999999998889***9***************************.****************** PP
ECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--..-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--S CS
START 85 issg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe.sssvvRaellpSgiliepksnghskvtwvehvdlkg 174
issg galqlm+aelq+lsplvp R++ f+R+++q+ +g+w++vdvS+d ++p++ + +v +++lpSg++i++++ng+skvtwveh+++++
Araip.PF1MQ 473 ISSGingtrnGALQLMQAELQVLSPLVPvREVNFLRFCKQHAEGVWAVVDVSIDTIREPSSaAPTFVSCRRLPSGCVIQDMPNGYSKVTWVEHAEYDE 570
************************************************************************************************** PP
SXXHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 175 rlphwllrslvksglaegaktwvatlqrqcek 206
+++h+l+r+lv+sg+ +ga++w atlqrqce+
Araip.PF1MQ 571 SQIHELYRPLVSSGMGFGAQRWIATLQRQCEC 602
******************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.04E-20 | 140 | 214 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 4.7E-21 | 148 | 214 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.394 | 154 | 214 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 9.9E-18 | 155 | 218 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.44E-18 | 156 | 214 | No hit | No description |
| Pfam | PF00046 | 1.5E-18 | 157 | 212 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 189 | 212 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 46.852 | 367 | 605 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.18E-35 | 369 | 602 | No hit | No description |
| CDD | cd08875 | 6.92E-123 | 371 | 601 | No hit | No description |
| SMART | SM00234 | 4.7E-45 | 376 | 602 | IPR002913 | START domain |
| Pfam | PF01852 | 4.1E-55 | 377 | 602 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 6.44E-24 | 630 | 854 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 861 aa Download sequence Send to blast |
MSFGGFLENK QGGGGARMVA EIPYSNGSSN HHHHHHHMEA ATAATIMASG AISHPHLAKS 60 MFNNSPALSL ALQTNMDGQG DVMNNNNRLL VGDNNNNNNN NTTNNNNYEG NMMSNNNGLR 120 RSREEEHESR SGSDNMDGAS GDDDLDAADI NNNKPRKKRY HRHTPQQIQE LEALFKECPH 180 PDEKQRLELS KRLCLETRQV KFWFQNRRTQ MKTQLERHEN TLLRQENDKL RAENMSLREA 240 MRNPICTNCG GPTMIGEISL EEQHLRIENA RLKDELDRVC ALAGKFLGRP IPSIAPHPPL 300 PNSSLELGVG NNNNSSNGGF VVGSTMPLGP DFVGMGISSN TLGVLSPTPP SRGIATTNGM 360 NSSNTSSGYE RSMVLELALG AMDELVKMAQ SGEPLWIRGM EMGGREVLNH EEYSRTVTPC 420 IGLRPNGFVS EASRETAIVI INSLPLVDTL MDSNRWSEMF PCMVARTSTT EVISSGINGT 480 RNGALQLMQA ELQVLSPLVP VREVNFLRFC KQHAEGVWAV VDVSIDTIRE PSSAAPTFVS 540 CRRLPSGCVI QDMPNGYSKV TWVEHAEYDE SQIHELYRPL VSSGMGFGAQ RWIATLQRQC 600 ECLAILMSSA VSPPEHSAIT SSGRRSMLKL AQRMTNNFCA GVCASTVHKW NKLNAGNFGE 660 DVRVMTRKSL DDPGEPPGIV LSAATSVWLP VSPQRLFDFL RDERLRSEWD ILSNGGPMQE 720 MAHIAKGQDH GNCVSLLRAS AMNSNQSSML ILQETCTDAS GSLVVYAPVD IPAMHLVMNG 780 GDSAYVALLP SGFAVAPDGS SGRNDHNTVS QPGGGSLLTV AFQILVNSLP TAKLTVESVE 840 TVNNLISCTV QKIKAALQSE G |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
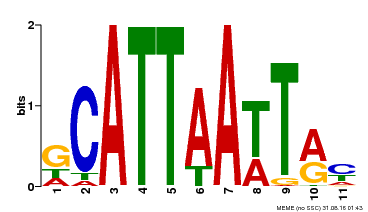
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Araip.PF1MQ |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_016198932.1 | 0.0 | homeobox-leucine zipper protein ANTHOCYANINLESS 2 isoform X2 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A444ZLZ2 | 0.0 | A0A444ZLZ2_ARAHY; Uncharacterized protein | ||||
| STRING | XP_007139955.1 | 0.0 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1192 | 34 | 106 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G61150.1 | 0.0 | homeodomain GLABROUS 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



