 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Araip.L2WGT | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Dalbergieae; Arachis
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1068aa MW: 120054 Da PI: 6.2798 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 181.3 | 1.1e-56 | 23 | 137 | 4 | 118 |
CG-1 4 ekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcywl 101
++rwl++ ei++iL n++ +++t+e+++rp+sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+hekLKvg+v+vl+cyYah+een++fqrr+yw+
Araip.L2WGT 23 AQHRWLRPAEICEILCNYRIFHITSEPPNRPPSGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHEKLKVGSVDVLHCYYAHGEENENFQRRSYWM 120
49************************************************************************************************ PP
CG-1 102 Leeelekivlvhylevk 118
Le ++ +iv+vhyl+vk
Araip.L2WGT 121 LEPDMMHIVFVHYLDVK 137
**************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 83.191 | 16 | 142 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 1.2E-81 | 19 | 137 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.2E-50 | 22 | 136 | IPR005559 | CG-1 DNA-binding domain |
| SuperFamily | SSF81296 | 6.86E-10 | 498 | 591 | IPR014756 | Immunoglobulin E-set |
| PROSITE profile | PS50297 | 18.006 | 688 | 797 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 8.1E-18 | 690 | 798 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 2.49E-17 | 694 | 797 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 2.04E-13 | 694 | 797 | No hit | No description |
| Pfam | PF12796 | 2.0E-6 | 710 | 793 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 9.965 | 738 | 770 | IPR002110 | Ankyrin repeat |
| SMART | SM00015 | 0.092 | 914 | 936 | IPR000048 | IQ motif, EF-hand binding site |
| SuperFamily | SSF52540 | 5.84E-7 | 914 | 961 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| PROSITE profile | PS50096 | 8.004 | 916 | 944 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0026 | 917 | 935 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0054 | 937 | 959 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.029 | 938 | 963 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.4E-4 | 940 | 959 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1068 aa Download sequence Send to blast |
MAERSSSFGL GPRLDLQQLQ FEAQHRWLRP AEICEILCNY RIFHITSEPP NRPPSGSLFL 60 FDRKVLRYFR KDGHNWRKKK DGKTVKEAHE KLKVGSVDVL HCYYAHGEEN ENFQRRSYWM 120 LEPDMMHIVF VHYLDVKVNK TNIGRNIDTD EAMSESQKGS SVLSGFPTSY NSMPSRSTDS 180 MSPTSTLTSL CEDADSEDIR QASSGMHSFG NPLGSGQAMD IIDTCSNRSY FMHPISGDHG 240 QSTISVTDYI PHVQGDKSRL GITAYIEDRK SHVMPLRDTI TEQSVGLYTP CSSISSGSRG 300 STLEQEKAVP ENLFGSKSGL NEESRGSQST QSNWQITFDE NTGPLPRWSL TESLSLEFES 360 DYSMALFGGE TDNAIPEICP DLFTSNGELK EQPVQQKFPK QSTDAQSQHA PRSDSEYELP 420 REDSAGYSLN AKHALLDGEE NLKKVDSFSR WITKELGEVD DLHLQSSPGI SWSTDECGHV 480 IDDTSLNPSL SQDQLFSISD FSPKWAYADS EIEVLIIGAF LKSQLEVEAC HWSCMFGEIE 540 VPXXXXXXXX XXXXNGIMCC QAPPRQVGRV PFYVTCSNRF ACSEVRDFEF REGFARNVDF 600 ADFFYSSTEM MLHLRLDELL SSKSVLSTNQ DFEGYMETRN LIFKLISLKE EEEYSHKEEA 660 TAGLSVSQHK LEEHVFHRKI KEKLYSWLLH KVTEGGKGPN VLDKDGQGVL HLVAALGYDW 720 AITPVITAGV NINFRDVNGW TALHWAASCG RERTVAVLVS MGAAAGALTD PSPAFPSGRT 780 AADLASSNGH KGISGFLGES LLTSHLASLT MDDPSEDGRN KTLGGKAVQT ASERSATPLL 840 YGDVSDTLCL KDSLSAVRNA TQAADRIHQV FRMQSFQRKQ LAQCEDDDEF GLSDQQVLSF 900 IASRACKSGQ GEGLVNAAAT QIQKKFRGWK KRKEFLIIRQ RVVKIQAHVR GHQVRKQYKL 960 IWSVGILEKV ILRWRRKGRG LRGFRPDTLN MVPNPPSNPS QEDDYDVLKE GRKQSEERFQ 1020 KALSRVKSMV QYPEARAQYR RVLNVVEDFR QTKVLYFLIY GLLPIAYD |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
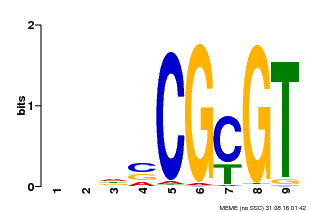
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Araip.L2WGT |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025639517.1 | 0.0 | calmodulin-binding transcription activator 2 isoform X1 | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | A0A445DMP5 | 0.0 | A0A445DMP5_ARAHY; Uncharacterized protein | ||||
| STRING | GLYMA05G24430.2 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF7406 | 26 | 42 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



