 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Neem_9045_f_2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Meliaceae; Azadirachta
|
||||||||
| Family | HSF | ||||||||
| Protein Properties | Length: 325aa MW: 36253.9 Da PI: 5.1282 | ||||||||
| Description | HSF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HSF_DNA-bind | 111.6 | 5.5e-35 | 10 | 102 | 2 | 103 |
HHHHHHHHHCTGGGTTTSEESSSSSEEEES-HHHHHHHTHHHHSTT--HHHHHHHHHHTTEEE---SSBTTTTXTTSEEEEESXXXXXXXXXXXXX CS
HSF_DNA-bind 2 FlkklyeiledeelkeliswsengnsfvvldeeefakkvLpkyFkhsnfaSFvRQLnmYgFkkvkdeekkskskekiweFkhksFkkgkkelleki 97
F++k+y++++d++++ li+w +nsf+v+++ +f++++Lp yFkh+nf+SFvRQLn+YgF+kv+ ++ weF++++F +g+k+ll++i
Neem_9045_f_2 10 FVMKTYQMVNDPTTDMLITWGIANNSFIVVEPLDFSQRILPAYFKHNNFSSFVRQLNTYGFRKVDPDR---------WEFANEWFLRGQKHLLKNI 96
9********************888*****************************************999.........******************* PP
XXXXXX CS
HSF_DNA-bind 98 krkkse 103
r+k++
Neem_9045_f_2 97 IRRKHS 102
999976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00415 | 5.9E-52 | 6 | 99 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Gene3D | G3DSA:1.10.10.10 | 4.2E-36 | 7 | 99 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SuperFamily | SSF46785 | 1.9E-31 | 7 | 99 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| Pfam | PF00447 | 7.0E-29 | 10 | 99 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 8.6E-18 | 10 | 33 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 8.6E-18 | 48 | 60 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PROSITE pattern | PS00434 | 0 | 49 | 73 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 8.6E-18 | 61 | 73 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 325 aa Download sequence Send to blast |
MESNNVIAPF VMKTYQMVND PTTDMLITWG IANNSFIVVE PLDFSQRILP AYFKHNNFSS 60 FVRQLNTYGF RKVDPDRWEF ANEWFLRGQK HLLKNIIRRK HSKSGAYLLK GEDLDDEEIV 120 MEIARLKEEQ KSLEEELQGM NKRLEATERR PEQMMAFLYK VAEDPDLLPR MVLEKERTKR 180 LGEKKRRLMI SSSQQPSSSS SGMAASSSIK SEEEEADGPL GLISSADSRF DTNFCQSSPS 240 PENNNDNNIS TGWFGQSRPI INYGCAAIPT PISAIPGTSG IANSMAVAAQ GNSISGQISY 300 FGEMATGGEP SPPPPYPFSL LGGGF |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5d5u_B | 4e-24 | 7 | 99 | 26 | 129 | Heat shock factor protein 1 |
| 5d5v_B | 4e-24 | 7 | 99 | 26 | 129 | Heat shock factor protein 1 |
| 5d5v_D | 4e-24 | 7 | 99 | 26 | 129 | Heat shock factor protein 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional regulator that specifically binds DNA sequence 5'-AGAAnnTTCT-3' known as heat shock promoter elements (HSE). | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00379 | DAP | Transfer from AT3G24520 | Download |
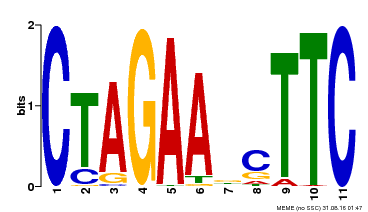
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002324418.2 | 1e-150 | heat stress transcription factor C-1 | ||||
| Swissprot | Q9LV52 | 1e-101 | HSFC1_ARATH; Heat stress transcription factor C-1 | ||||
| TrEMBL | B9ILM2 | 1e-148 | B9ILM2_POPTR; Uncharacterized protein | ||||
| STRING | cassava4.1_010201m | 1e-148 | (Manihot esculenta) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM6775 | 27 | 43 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G24520.1 | 6e-92 | heat shock transcription factor C1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



