 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Neem_7557_f_9 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Meliaceae; Azadirachta
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 352aa MW: 39692.6 Da PI: 8.906 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 92.2 | 4.5e-29 | 62 | 118 | 2 | 54 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQ....kYR 54
prlrWtp+LH +Fv+ave+LGG+e+AtPk +l+lm++kgL+++hvkSHLQ +YR
Neem_7557_f_9 62 PRLRWTPDLHLCFVQAVEKLGGQERATPKLVLQLMNIKGLSIAHVKSHLQarcsMYR 118
8*************************************************5555566 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 7.7E-27 | 58 | 119 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 10.537 | 58 | 122 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 6.63E-13 | 60 | 116 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.0E-20 | 62 | 112 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 2.9E-8 | 63 | 111 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 352 aa Download sequence Send to blast |
MTKKREISNG LEMNLFGRDE DEKGDDESKT KTSASSSNSV VEDGEKKASS SGVRPYVRSK 60 VPRLRWTPDL HLCFVQAVEK LGGQERATPK LVLQLMNIKG LSIAHVKSHL QARCSMYRSK 120 KIDDHGQVIN SRGHLMRSSD HFSHNLWHHS MLRGINQRIG SNFSDASWIG HGNSMLRPHM 180 NIGSRACFGE PLLARSIYGE IGSNIRNEAL YTRRNLAFSE QRIGRTSERH EDCLSYNCDA 240 FQTQSSSTVP SLWHGKERTK CCNYTPNPAN PNSKWMIDGL EEQSTGKRKA LEHDDLDLSL 300 SLGSKVRKEE VRRKLLGDEE EVESILLLSL SSSLTREICS TNLQRHSEAN NS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 3e-16 | 63 | 118 | 4 | 60 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 3e-16 | 63 | 118 | 4 | 60 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 3e-16 | 63 | 118 | 4 | 60 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 3e-16 | 63 | 118 | 4 | 60 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 3e-16 | 63 | 118 | 4 | 60 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 3e-16 | 63 | 118 | 4 | 60 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 3e-16 | 63 | 118 | 4 | 60 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 3e-16 | 63 | 118 | 4 | 60 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
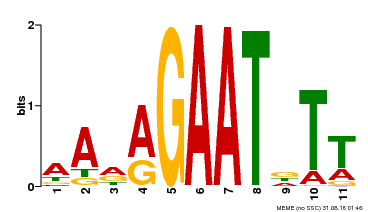
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00297 | DAP | Transfer from AT2G38300 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015388811.1 | 1e-137 | uncharacterized protein LOC107178351 | ||||
| TrEMBL | V4SHP8 | 1e-136 | V4SHP8_9ROSI; Uncharacterized protein | ||||
| STRING | XP_006425115.1 | 1e-137 | (Citrus clementina) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM363 | 28 | 184 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G38300.1 | 6e-44 | G2-like family protein | ||||



