 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Neem_7267_f_1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Meliaceae; Azadirachta
|
||||||||
| Family | E2F/DP | ||||||||
| Protein Properties | Length: 592aa MW: 65777.8 Da PI: 5.1949 | ||||||||
| Description | E2F/DP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | E2F_TDP | 82.7 | 3e-26 | 252 | 332 | 1 | 71 |
E2F_TDP 1 rkeksLrlltqkflkllekseegivtlnevakeLvse..............dvknkrRRiYDilNVLealnliekkekneirwkg 71
r+++sL+llt+kf++l++++e+gi++ln++a++L e +v ++RRiYDi+NVLe+++liekk kn+irwkg
Neem_7267_f_1 252 RYDSSLGLLTKKFINLIKHAEDGILDLNKAAETL--EviiwpltsytveyfNV--QKRRIYDITNVLEGIGLIEKKLKNRIRWKG 332
6899******************************..779**********9999..****************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.10 | 8.1E-27 | 249 | 334 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SuperFamily | SSF46785 | 7.35E-14 | 252 | 330 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SMART | SM01372 | 2.1E-29 | 252 | 332 | IPR003316 | E2F/DP family, winged-helix DNA-binding domain |
| Pfam | PF02319 | 4.4E-23 | 254 | 332 | IPR003316 | E2F/DP family, winged-helix DNA-binding domain |
| CDD | cd14660 | 1.20E-44 | 346 | 449 | No hit | No description |
| SuperFamily | SSF144074 | 3.4E-31 | 346 | 449 | No hit | No description |
| Pfam | PF16421 | 3.9E-31 | 348 | 447 | IPR032198 | E2F transcription factor, CC-MB domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005667 | Cellular Component | transcription factor complex | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 592 aa Download sequence Send to blast |
MSGTTRASSC PEPSLPAGPI VLPMRHHLAP VTTKPPFVPS DEYHRLSTTR GAADQQPEVI 60 FVRSPQLRRK TGGNGNDVES DTSMSTKANI RSKAAKRNRS TSQTPLSSTG SRRCDSYLGS 120 NPENLIIYAL KLSKKAGIEK LSIEECRVDV KIRKETLILI KAPKETSLEV PDPDEQLKRK 180 TGMDDNEVES SQWVGSPGLT NISNGSFQTP VSAKGGRVNN RSKATKGNRS GPQTPVSNAG 240 SPATLTPAGS CRYDSSLGLL TKKFINLIKH AEDGILDLNK AAETLEVIIW PLTSYTVEYF 300 NVQKRRIYDI TNVLEGIGLI EKKLKNRIRW KGLDNSTPGE VDGDASLLQA EIENLTMEER 360 RVDDQIREMQ ERLRDLSENE NNRKWLFVTE EDIKTVPCFL NETLIAIKAP HGTTLEVPDP 420 DEAVDYPQRR YRIILRSTMG PIDVYLVSQF EEKFEEMNNV VPAESFSIAC NSVSKENQER 480 GIINVESTRN EIEAEAQHAH QICSDINSSE DFGGGMMKIV PSDVDNDADY WLLSDADVSI 540 TDMWKTDSGV EWNGVNMLHP DFGMSDVCSP RPQTPPSRIS EVPSTDFNST HR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1cf7_A | 6e-22 | 247 | 333 | 2 | 74 | PROTEIN (TRANSCRIPTION FACTOR E2F-4) |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds DNA cooperatively with DP proteins through the E2 recognition site, 5'-TTTC[CG]CGC-3' found in the promoter region of a number of genes whose products are involved in cell cycle regulation or in DNA replication. The binding of retinoblastoma-related proteins represses transactivation. Regulates gene expression both positively and negatively. Activates the expression of E2FB. Involved in the control of cell-cycle progression from G1 to S phase. Stimulates cell proliferation and delays differentiation. {ECO:0000269|PubMed:11669580, ECO:0000269|PubMed:11786543, ECO:0000269|PubMed:11862494, ECO:0000269|PubMed:11889041, ECO:0000269|PubMed:11891240, ECO:0000269|PubMed:12913157, ECO:0000269|PubMed:15377755, ECO:0000269|PubMed:16514015, ECO:0000269|PubMed:19662336}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00661 | PBM | Transfer from Pp3c1_6880 | Download |
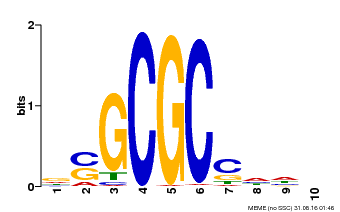
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006481571.1 | 0.0 | transcription factor E2FA isoform X3 | ||||
| Refseq | XP_024038059.1 | 0.0 | transcription factor E2FA isoform X3 | ||||
| Swissprot | Q9FNY0 | 1e-151 | E2FA_ARATH; Transcription factor E2FA | ||||
| TrEMBL | V4SRJ4 | 0.0 | V4SRJ4_9ROSI; Uncharacterized protein | ||||
| STRING | XP_006481568.1 | 0.0 | (Citrus sinensis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM808 | 28 | 121 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G36010.1 | 1e-155 | E2F transcription factor 3 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



