 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Neem_606_f_9 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Meliaceae; Azadirachta
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 670aa MW: 74975.8 Da PI: 5.8986 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 26 | 2e-08 | 200 | 228 | 1 | 29 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtP 29
k+r++W+ +LH++Fv+av+q+G +k P
Neem_606_f_9 200 KARVVWSVDLHQKFVKAVNQIGFDKKMLP 228
68*******************88777777 PP
| |||||||
| 2 | G2-like | 25.9 | 2.2e-08 | 250 | 273 | 28 | 51 |
G2-like 28 tPktilelmkvkgLtlehvkSHLQ 51
Pk+il+lm+v+ Lt+e+v+SHLQ
Neem_606_f_9 250 GPKKILDLMNVPWLTRENVASHLQ 273
6*********************** PP
| |||||||
| 3 | G2-like | 37.9 | 4.1e-12 | 296 | 323 | 28 | 55 |
G2-like 28 tPktilelmkvkgLtlehvkSHLQkYRl 55
Pk+il+lm+v+ Lt+e+v+SHLQkYRl
Neem_606_f_9 296 GPKKILDLMNVPWLTRENVASHLQKYRL 323
5**************************8 PP
| |||||||
| 4 | Response_reg | 80.7 | 4.9e-27 | 20 | 128 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTTSEEEEEESTTTHHHHHHHHHTTES CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepklpiivvtahgeeedalealkaGak 95
vl+vdD+p+ +++l+++l+k y ev+++ +++al ll+e++ +D+++ D++mp+mdG++ll++ e +lp+i+++ ge + + + ++ Ga
Neem_606_f_9 20 VLVVDDDPTWLKILEKMLKKCSY-EVTTCGLARDALSLLRERKdgYDIVISDVNMPDMDGFKLLEHVGLEM-DLPVIMMSVDGETSRVMKGVQHGAC 114
89*********************.***************999999**********************6644.8************************ PP
EEEESS--HHHHHH CS
Response_reg 96 dflsKpfdpeelvk 109
d+l Kp+ ++el++
Neem_606_f_9 115 DYLLKPIRMKELRN 128
***********987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:3.40.50.2300 | 2.6E-44 | 17 | 161 | No hit | No description |
| SuperFamily | SSF52172 | 2.28E-35 | 17 | 142 | IPR011006 | CheY-like superfamily |
| SMART | SM00448 | 6.9E-32 | 18 | 130 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 43.143 | 19 | 134 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 4.1E-24 | 20 | 129 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 1.40E-29 | 21 | 134 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 4.9E-20 | 198 | 273 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 4.3E-9 | 198 | 274 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.5E-7 | 248 | 274 | IPR006447 | Myb domain, plants |
| SuperFamily | SSF46689 | 1.16E-6 | 293 | 327 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 3.5E-10 | 295 | 323 | IPR006447 | Myb domain, plants |
| Gene3D | G3DSA:1.10.10.60 | 4.2E-13 | 295 | 328 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000160 | Biological Process | phosphorelay signal transduction system | ||||
| GO:0009735 | Biological Process | response to cytokinin | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 670 aa Download sequence Send to blast |
MENGFSSPRN DAFNPAGLRV LVVDDDPTWL KILEKMLKKC SYEVTTCGLA RDALSLLRER 60 KDGYDIVISD VNMPDMDGFK LLEHVGLEMD LPVIMMSVDG ETSRVMKGVQ HGACDYLLKP 120 IRMKELRNIW QHVFRKRIHE VRDIENIEGF ESIQMTRSGS DQSDDGHFLP GEDLTFARKR 180 KDAESKHDDK DSGDASSTKK ARVVWSVDLH QKFVKAVNQI GFDKKMLPAT WSLSCFHRAM 240 TCNMCLAEVG PKKILDLMNV PWLTRENVAS HLQVDTLRFC CQVKLFIYFP SFREVGPKKI 300 LDLMNVPWLT RENVASHLQK YRLYLSRLQK ENELKSAIGG IKQTDSPSKD SANSFGLQNS 360 INLHHHNDVS SGSYGFSGSN SLVQNVNPKG LEGDLKGIVT IPVAESKRPL TVDIPNQRNI 420 RSSRNSQIGF DCSFAAGESE VSFTAFDSTI PSQYPWSDVP ETQFKPEHKP LQLEDGYSQL 480 SLPAPGHHIQ NDRLQSCPSI SPGPSVADRD NNLKPLDAMY KSNYASAIPV GSAVDSFPVQ 540 TKSHMVNHQS FEPIPTAISS IKSQGLNLSC ISDLESAQRN INWGFGSSLA ALDDDLQFCW 600 LQGDCLAMNL GLQDINCSGY NDPGLISEVP FHLYDTMRFD YDHYYDPTEY SVIDQDGCIA 660 VMGDNAQGQK |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00010 | PBM | Transfer from AT1G67710 | Download |
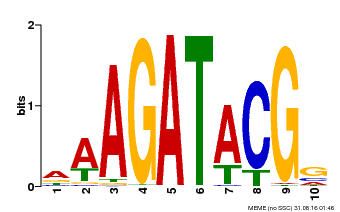
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006484016.1 | 0.0 | two-component response regulator ARR11 isoform X1 | ||||
| Refseq | XP_006484017.1 | 0.0 | two-component response regulator ARR11 isoform X2 | ||||
| TrEMBL | A0A2H5NV41 | 0.0 | A0A2H5NV41_CITUN; Uncharacterized protein | ||||
| STRING | XP_006484017.1 | 0.0 | (Citrus sinensis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM8066 | 27 | 40 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G67710.1 | 1e-131 | response regulator 11 | ||||



