 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Neem_544_f_1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Meliaceae; Azadirachta
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 819aa MW: 89250.7 Da PI: 6.3569 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 65.4 | 7.9e-21 | 109 | 164 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+++l L++rqVk+WFqNrR+++k
Neem_544_f_1 109 KKRYHRHTPQQIQELEALFKECPHPDEKQRLELSRRLCLETRQVKFWFQNRRTQMK 164
688999***********************************************999 PP
| |||||||
| 2 | START | 209.6 | 1.2e-65 | 318 | 542 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEEEEE CS
START 1 elaeeaaqelvkkalaeepgWvkss....esengdevlqkfeeskv.....dsgealrasgvvdmvlallveellddkeqWdetla....kaetlev 84
ela++a++elvk+a+ +ep+W +s e +n++e++++f++ + + +ea+r+sg+v+ ++ lve+l+d + +W e+++ + +t++v
Neem_544_f_1 318 ELALAAMDELVKMAQTDEPLWIRSLeggrEVLNREEYMRTFTPCIGmkptgFVTEASRESGMVIINSLALVETLMDPN-RWAEMFPcmiaRTSTTDV 413
5899**************************************99989*******************************.****************** PP
ECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--.-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--S CS
START 85 issg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppesssvvRaellpSgiliepksnghskvtwvehvdlkg 174
is+g galqlm aelq+lsplvp R++ f+R+++q+ + +w++vdvS+d ++ ++ + +v +++lpSg+++++++ng+skvtwveh+++++
Neem_544_f_1 414 ISNGmggtrnGALQLMHAELQVLSPLVPvREVNFLRFCKQHAESVWAVVDVSIDTIRETSSAPTFVNCRRLPSGCVVQDMPNGYSKVTWVEHAEYDE 510
**********************************************************999************************************ PP
SXXHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 175 rlphwllrslvksglaegaktwvatlqrqcek 206
+++h+l+++l++sg+ +ga++wvatlqrqce+
Neem_544_f_1 511 SQVHQLYKPLISSGMGFGAQRWVATLQRQCEC 542
******************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.2E-20 | 92 | 166 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 1.4E-21 | 95 | 166 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.216 | 106 | 166 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.7E-17 | 107 | 170 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 2.02E-18 | 108 | 166 | No hit | No description |
| Pfam | PF00046 | 2.2E-18 | 109 | 164 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 141 | 164 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 44.353 | 309 | 545 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 2.7E-35 | 311 | 542 | No hit | No description |
| CDD | cd08875 | 8.12E-129 | 313 | 541 | No hit | No description |
| Pfam | PF01852 | 1.9E-57 | 318 | 542 | IPR002913 | START domain |
| SMART | SM00234 | 1.9E-48 | 318 | 542 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.51E-21 | 570 | 812 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 819 aa Download sequence Send to blast |
MSFGGFLDNS SGGGGARIVA DISYSNNMPT TTLAQPRLVT PSLTKSMFNS PGLSLALIIK 60 VTCLEWGENF ESVAGRRSRE EEHESRSGSE NMDGASGDDQ DAADNPPRKK RYHRHTPQQI 120 QELEALFKEC PHPDEKQRLE LSRRLCLETR QVKFWFQNRR TQMKTQLERH ENSLLRQEND 180 KLRAENMSIR DAMRNPICTN CGGPAIIGDI SLEEQHLRIE NARLKDELDR VCALAGKFLG 240 RPISSLATSM APPMPNSSLE LGVGSNGFGG LSPVTTTLPL GPDFGGGISN PLPVMPPNRS 300 TAGVTGLDRS IERSMFLELA LAAMDELVKM AQTDEPLWIR SLEGGREVLN REEYMRTFTP 360 CIGMKPTGFV TEASRESGMV IINSLALVET LMDPNRWAEM FPCMIARTST TDVISNGMGG 420 TRNGALQLMH AELQVLSPLV PVREVNFLRF CKQHAESVWA VVDVSIDTIR ETSSAPTFVN 480 CRRLPSGCVV QDMPNGYSKV TWVEHAEYDE SQVHQLYKPL ISSGMGFGAQ RWVATLQRQC 540 ECLAILMSST VSARDHTAIT AGGRRSMLKL AQRMTDNFCA GVCASTVHKW NKLNAGNVDE 600 DVRVMTRKSV DDPGEPPGIV LSAATSVWLP VSPQRLFDFL RDERLRSEWD ILSNGGPMQE 660 MAHIAKGQDH GNCVSLLRAS FDSRNAVKAM NANQSSMLIL QETCIDAAGS LVVYAPVDIP 720 AMHVVMNGGD SAYVALLPSG FAIVPDGPGS RGPMANGPTS GNGGNGGSPR VSGSLLTVAF 780 QILVNSLPTA KLTVESVETV NNLISCTVQK IKAALQCES |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
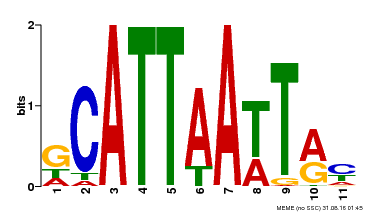
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AM459307 | 1e-39 | AM459307.2 Vitis vinifera contig VV78X101111.6, whole genome shotgun sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010661562.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ANTHOCYANINLESS 2 isoform X2 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A061DTK7 | 0.0 | A0A061DTK7_THECC; HD domain class transcription factor isoform 2 | ||||
| STRING | VIT_15s0048g02000.t01 | 0.0 | (Vitis vinifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1128 | 27 | 105 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



