 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Neem_3898_f_3 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Meliaceae; Azadirachta
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 620aa MW: 70316.6 Da PI: 6.5801 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 86.2 | 5.1e-27 | 90 | 192 | 1 | 91 |
FAR1 1 kfYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkk............tekerrtraetrtgCkaklkvkkekdgkwevtklele 84
+fY+eYA+++GF++ +++s++sk+++e+++++f Cs++g+++e +k+ +e++ +r+ +t+Cka+++vk++ dgkw+++++++e
Neem_3898_f_3 90 SFYQEYARSMGFNTAIQNSRRSKTSREFIDAKFACSRYGTKREYDKSynrprarqskqdQENATGRRSCAKTDCKASMHVKRRPDGKWVIHSFVKE 185
5******************************************99999****99887655555669999*************************** PP
FAR1 85 HnHelap 91
HnHel p
Neem_3898_f_3 186 HNHELLP 192
****975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 7.1E-25 | 90 | 192 | IPR004330 | FAR1 DNA binding domain |
| PROSITE profile | PS50966 | 9.26 | 341 | 377 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 1.7E-5 | 351 | 374 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 3.2E-7 | 352 | 379 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010218 | Biological Process | response to far red light | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 620 aa Download sequence Send to blast |
MDIDLRLPSG EQAKEEEEQN GIDNMLDGEE KLALHNGDVE TGNIAVVDEV RAEDGGGVNS 60 PAEDMVVFKE DTNLEPLSGM EFESHGEAYS FYQEYARSMG FNTAIQNSRR SKTSREFIDA 120 KFACSRYGTK REYDKSYNRP RARQSKQDQE NATGRRSCAK TDCKASMHVK RRPDGKWVIH 180 SFVKEHNHEL LPAQAVSEQT RKMYAAMARQ FAEYKSVVGL KNDPKNPFDK GRNLALELGD 240 AKILLDFFTQ MQNMNANFFY AVDLDRYEEE AKADSDTWNK QPALKSPSPF EKSVSAIYTH 300 AVFKKFQVEV LGAVACHPKQ ESQNEANVMF RVQDFEKNQD FIVMWNQMKS EVFCVCRLFE 360 YKGYLCRHAL IVLQIRGLSA IPSQYILKRW TRDAKSRQLL GEESDQIQSR VQRYNDLCQR 420 AMKLSEEGSL SQESYGVALR ALDEALANCS SVNTSNKSLV EAVTSPTHGL ICIEEDNQSR 480 SMSKTNKRKN PTKKRKVNSE QEVMTVGAQD SLQQMEKLNS RAVTSLDGYY GTQQSVQGMV 540 QLNLMAPTRD NYYSNQQTIQ GLGQLNSIAP SHDGYYSAQQ GMHGLGQMDF FRTPTSFTYG 600 IRDDPNVRTA QLQDDASRHA |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
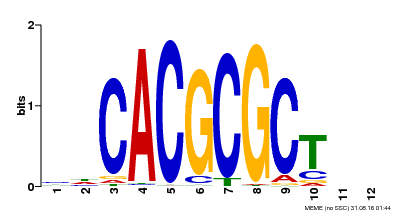
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. When associated with PHYA, protects it from being recognized and degraded by the COP1/SPA complex. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:17012604, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:18715961, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00078 | ChIP-seq | Transfer from AT3G22170 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. Up-regulated by white light. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006481370.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X1 | ||||
| Refseq | XP_024037340.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| Refseq | XP_024037341.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| Refseq | XP_024037342.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| Refseq | XP_024037343.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| Refseq | XP_024955384.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X1 | ||||
| Refseq | XP_024955385.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X1 | ||||
| Refseq | XP_024955386.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X1 | ||||
| Refseq | XP_024955387.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X1 | ||||
| Swissprot | Q9LIE5 | 1e-165 | FHY3_ARATH; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| TrEMBL | A0A3N6QE62 | 0.0 | A0A3N6QE62_BRACR; Uncharacterized protein | ||||
| STRING | XP_006481370.1 | 0.0 | (Citrus sinensis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM118 | 26 | 253 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G22170.2 | 1e-165 | far-red elongated hypocotyls 3 | ||||



