 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Neem_2491_f_1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Meliaceae; Azadirachta
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 372aa MW: 40917.1 Da PI: 8.6119 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 100.8 | 8.8e-32 | 201 | 256 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r++W+peLH+rF++a++qLGGs+ AtPk+i+e mkv+gLt+++vkSHLQkYRl+
Neem_2491_f_1 201 KQRRCWSPELHRRFLHALQQLGGSHAATPKQIREQMKVDGLTNDEVKSHLQKYRLH 256
79****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 4.1E-28 | 198 | 259 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 1.25E-17 | 198 | 259 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 14.887 | 198 | 258 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 6.4E-27 | 201 | 256 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.4E-7 | 203 | 254 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 372 aa Download sequence Send to blast |
MDYAEKMQRC HEYVEALEEE KRKIQVFQRE LPLSIEACRR ELSGTTTEYM HGQSECSEQT 60 SSDVPVLEEF MPIKRSSSSE DNDEQHSHKN TTSNREKNSD KKKSDWLRSV QLWNQSPDPL 120 PKEDVPRKTT VVEVKRNNGG AFQPFQREKS VGKTEQSAGK VNSSLPAPAA SSTTETVTKG 180 GGGGGSGSSK KDEKEGQTQR KQRRCWSPEL HRRFLHALQQ LGGSHAATPK QIREQMKVDG 240 LTNDEVKSHL QKYRLHTRRP TPSIHGNSNP QAPQFVVVGG IWVQPPEYAT VTAKSSAEAS 300 TMTAANGIYA PLAAPPPALT QSSSPMQRPQ QAQSDHLQSE ERGSHSEGGG AHHSNSPATS 360 SSTHTTTASP AF |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in phosphate signaling in roots. {ECO:0000250|UniProtKB:Q9FX67}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00160 | DAP | Transfer from AT1G25550 | Download |
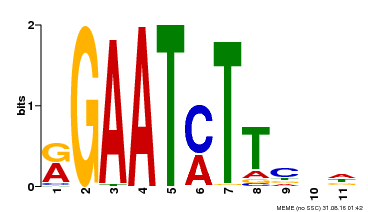
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006437730.1 | 1e-173 | transcription factor HHO3 | ||||
| Swissprot | Q9FPE8 | 1e-111 | HHO3_ARATH; Transcription factor HHO3 | ||||
| TrEMBL | V4TH64 | 1e-171 | V4TH64_9ROSI; Uncharacterized protein | ||||
| STRING | XP_006437730.1 | 1e-172 | (Citrus clementina) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3491 | 28 | 63 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G25550.1 | 1e-103 | G2-like family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



