 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Neem_2214_f_1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Meliaceae; Azadirachta
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 834aa MW: 91984.6 Da PI: 6.5047 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 58.4 | 1.2e-18 | 17 | 75 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++++++ps +r++L +++ +++ +q+kvWFqNrR +ek+
Neem_2214_f_1 17 KYVRYTPEQVEALERLYHECPKPSSIRRQQLIRECpilsNIEPKQIKVWFQNRRCREKQ 75
5679*****************************************************97 PP
| |||||||
| 2 | START | 177.7 | 6.8e-56 | 144 | 352 | 2 | 205 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEECTT..EEEEEEE CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetlevissg..galqlmv 95
+aee+++e+++ka+ ++ Wv+++ +++g++++ +++ s+++ g a+ra+g+v +++ v+e+l+d++ W ++++++++l+v+ ++ g+++l +
Neem_2214_f_1 144 IAEETLAEFLSKATGTAVEWVQMPGMKPGPDSVGIVAISHSCAGVAARACGLVGLEPT-RVAEILKDRPSWFRDCRAVDVLNVLPTAngGTIELLY 238
7899******************************************************.8888888888****************9999******* PP
EXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHHHHHHHHHH CS
START 96 aelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwvehvdlkgrlphwllrslvks 187
++l+al++l+p Rdf+ +Ry+ l++g++v++++S+ ++q+ p+ +++vRae lpSg+li+p+++g+s +++v+h+dl+ ++++++lr+l++s
Neem_2214_f_1 239 MQLYALTTLAPaRDFWLLRYTSVLEDGSLVVCERSLKNTQNGPSmppVQHFVRAEILPSGYLIRPCEGGGSIIHIVDHMDLEPWSVPEVLRPLYES 334
******************************************9988899*********************************************** PP
HHHHHHHHHHHHTXXXXX CS
START 188 glaegaktwvatlqrqce 205
+++ ++kt++a+l+++++
Neem_2214_f_1 335 STVLAQKTTMAALRQLRQ 352
**************9876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 15.548 | 12 | 76 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.2E-15 | 14 | 80 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 4.28E-17 | 16 | 80 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 9.24E-17 | 17 | 77 | No hit | No description |
| Pfam | PF00046 | 2.9E-16 | 18 | 75 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 1.6E-18 | 19 | 75 | IPR009057 | Homeodomain-like |
| CDD | cd14686 | 3.31E-6 | 69 | 106 | No hit | No description |
| PROSITE profile | PS50848 | 25.78 | 134 | 362 | IPR002913 | START domain |
| CDD | cd08875 | 3.77E-76 | 138 | 354 | No hit | No description |
| SuperFamily | SSF55961 | 4.81E-37 | 143 | 355 | No hit | No description |
| Gene3D | G3DSA:3.30.530.20 | 3.5E-23 | 143 | 350 | IPR023393 | START-like domain |
| SMART | SM00234 | 9.8E-41 | 143 | 353 | IPR002913 | START domain |
| Pfam | PF01852 | 4.9E-53 | 144 | 352 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.1E-5 | 389 | 478 | No hit | No description |
| SuperFamily | SSF55961 | 1.1E-5 | 510 | 588 | No hit | No description |
| Pfam | PF08670 | 1.9E-51 | 691 | 833 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 834 aa Download sequence Send to blast |
MAMSCKDGKP GSFDNGKYVR YTPEQVEALE RLYHECPKPS SIRRQQLIRE CPILSNIEPK 60 QIKVWFQNRR CREKQRKEAS RLQAVNRKLT AMNKLLMEEN DRLQKQTTLA TKDTSCESVV 120 TSGQHHLTPQ HPPRDASPAG LLSIAEETLA EFLSKATGTA VEWVQMPGMK PGPDSVGIVA 180 ISHSCAGVAA RACGLVGLEP TRVAEILKDR PSWFRDCRAV DVLNVLPTAN GGTIELLYMQ 240 LYALTTLAPA RDFWLLRYTS VLEDGSLVVC ERSLKNTQNG PSMPPVQHFV RAEILPSGYL 300 IRPCEGGGSI IHIVDHMDLE PWSVPEVLRP LYESSTVLAQ KTTMAALRQL RQIAQEVSQS 360 SVTGWGRRPA ALRALSQRLS RQMLHYRGFN EAVNGFTDEG WTVMGNDGMD DVTILVNSSP 420 EKLMGLNLSF GNGFPTVSNA VLCAKASMLL QNVPPAILLR FLREHRSEWA DNNIDVYSAA 480 AIKVGPCTLP GSRVGSFGSQ VILPLAHTIE HEEFMEVIKL EGVGHSPEDT MMPRDMFLLQ 540 LCSGMDENAV GTCAELIFAP IDASFADDAP VLPSGFRIIP LDSGKMLLNT IQEASSPNRT 600 LDLASALEIG PAGNRASNDY SSNSSCMRSV MTIAFEFAFE SHMQEHVATM ARQYVRSIIS 660 SVQRVALALS PSHLSSQAGL RTPLGTPEAQ TLARWICQSY RCYLGVELLK SSSEGSESIL 720 KTLWHHTDAV MCCSLKALPV FTFANQAGLD LLETTLVALQ DITLEKMFDD HGRKALFSEF 780 PQIMQQGFAC LQGGICLSSM GRPVSYERAV AWKVLNEEEA AHCICFMFIN WSFV |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
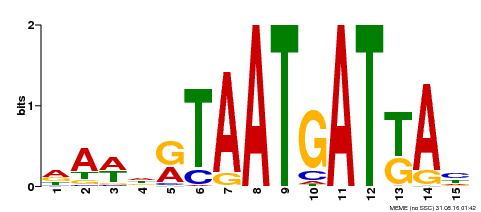
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006426893.1 | 0.0 | homeobox-leucine zipper protein ATHB-15 | ||||
| Refseq | XP_006465680.1 | 0.0 | homeobox-leucine zipper protein ATHB-15 | ||||
| Refseq | XP_015386901.1 | 0.0 | homeobox-leucine zipper protein ATHB-15 | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | V4RYP0 | 0.0 | V4RYP0_9ROSI; Uncharacterized protein | ||||
| STRING | XP_006465679.1 | 0.0 | (Citrus sinensis) | ||||
| STRING | XP_006426892.1 | 0.0 | (Citrus clementina) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2022 | 26 | 78 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.2 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



