 |
Plant Transcription
Factor Database
|
Transcription Factor Information
|
Basic
Information? help
Back to Top |
| TF ID |
Neem_18380_f_1 |
| Organism |
|
| Taxonomic ID |
|
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Meliaceae; Azadirachta
|
| Family |
NF-X1 |
| Protein Properties |
Length: 250aa MW: 27594.5 Da PI: 8.4455 |
| Description |
NF-X1 family protein |
| Gene Model |
|
| Signature Domain? help Back to Top |
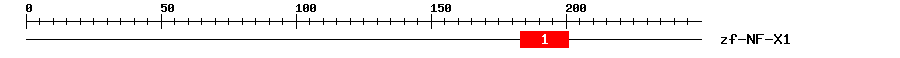 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | zf-NF-X1 | 18.1 | 5.8e-06 | 183 | 201 | 1 | 20 |
zf-NF-X1 1 CGkHkCqklCHeGpCppCpq 20
CG H C lCH+GpCp+Cp+
Neem_18380_f_1 183 CG-HFCLLLCHPGPCPSCPK 201
**.***************96 PP
|
| Gene Ontology ? help Back to Top |
| GO Term |
GO Category |
GO Description |
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated |
| GO:0005634 | Cellular Component | nucleus |
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding |
| GO:0005515 | Molecular Function | protein binding |
| GO:0008270 | Molecular Function | zinc ion binding |
| Sequence ? help Back to Top |
| Protein Sequence Length: 250 aa
Download sequence Send
to blast |
MTSTANYQQL LSDSDSDSDN GRPESFPDLS SSILKPYLES TNQSLLTASP SDLSKIRSFL 60
TSSSSGALSC LICLERIRPS DPTWHCSSLC YSIFHLICIQ SWARQASDLS ASFAASRLPI 120
SAAKAAESST WNCPKCRSQY SKSEIPKKYV CFCGKLEDPY SEDPWILPHS CGEVCNRPLQ 180
HNCGHFCLLL CHPGPCPSCP KLVKSSCFCG KIVRLMTVQK SATTGRARRV KRGRYTNADV 240
AEFKKRESVV
|
| Functional Description ? help
Back to Top |
| Source |
Description |
| UniProt | Probable transcriptional regulator. May mediate E2- or E3-dependent ubiquitination. Required to gate light sensitivity during the night. Regulates the speed of the clock by acting in the feedback loop between CCA1, LHY and APRR1/TOC1. Promotes the expression of CCA1 at night but not by days. This activational effect is enhanced by interaction with ADO1/ZTL. Association with ADO1/ZTL is not leading to the degradation of NFXL2. Confers sensitivity to osmotic stress such as high salinity. Prevents H(2)O(2) production and abscisic acid accumulation. Part of a regulatory network that integrates the biosynthesis and action of abscisic acid, reactive oxygen species and cuticle components. {ECO:0000269|PubMed:16905136, ECO:0000269|PubMed:21300918, ECO:0000269|PubMed:21429190, ECO:0000269|PubMed:22073231, ECO:0000269|PubMed:22516817}. |
| Regulation -- Description ? help
Back to Top |
| Source |
Description |
| UniProt | INDUCTION: Circadian-regulation with a peak of expression at or before dawn. Not regulated by biotic and abiotic stresses, by light and nutrient conditions or upon treatment with elicitors, chemicals, abscisic acid or phytohormones. {ECO:0000269|PubMed:21300918, ECO:0000269|PubMed:22073231}. |
| Best hit in Arabidopsis thaliana ? help
Back to Top |
| Hit ID |
E-value |
Description |
| AT5G05660.1 | 2e-62 | sequence-specific DNA binding transcription factors;zinc ion binding;sequence-specific DNA binding transcription factors |




